Distinct molecular mechanisms underlying clinically relevant subtypes of breast cancer: gene expression analyses across three different platforms
- PMID: 16729877
- PMCID: PMC1489944
- DOI: 10.1186/1471-2164-7-127
Distinct molecular mechanisms underlying clinically relevant subtypes of breast cancer: gene expression analyses across three different platforms
Abstract
Background: Gene expression profiling has been used to define molecular phenotypes of complex diseases such as breast cancer. The luminal A and basal-like subtypes have been repeatedly identified and validated as the two main subtypes out of a total of five molecular subtypes of breast cancer. These two are associated with distinctly different gene expression patterns and more importantly, a significant difference in clinical outcome. To further validate and more thoroughly characterize these two subtypes at the molecular level in tumors at an early stage, we report a gene expression profiling study using three different DNA microarray platforms.
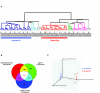
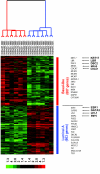
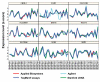
Results: Expression data from 20 tumor biopsies of early stage breast carcinomas were generated on three different DNA microarray platforms; Applied Biosystems Human Genome Survey Microarrays, Stanford cDNA Microarrays and Agilent's Whole Human Genome Oligo Microarrays, and the resulting gene expression patterns were analyzed. Both unsupervised and supervised analyses identified the different clinically relevant subtypes of breast tumours, and the results were consistent across all three platforms. Gene classification and biological pathway analyses of the genes differentially expressed between the two main subtypes revealed different molecular mechanisms descriptive of the two expression-based subtypes: Signature genes of the luminal A subtype were over-represented by genes involved in fatty acid metabolism and steroid hormone-mediated signaling pathways, in particular estrogen receptor signaling, while signature genes of the basal-like subtype were over-represented by genes involved in cell proliferation and differentiation, p21-mediated pathway, and G1-S checkpoint of cell cycle-signaling pathways. A minimal set of 54 genes that best discriminated the two subtypes was identified using the combined data sets generated from the three different array platforms. These predictor genes were further verified by TaqMan Gene Expression assays.
Conclusion: We have identified and validated the two main previously defined clinically relevant subtypes, luminal A and basal-like, in a small set of early stage breast carcinomas. Signature genes characterizing these two subtypes revealed that distinct molecular mechanisms might have been pre-programmed at an early stage in different subtypes of the disease. Our results provide further evidence that these breast tumor subtypes represent biologically distinct disease entities and may require different therapeutic strategies. Finally, validated by multiple gene expression platforms, including quantitative PCR, the set of 54 predictor genes identified in this study may define potential prognostic molecular markers for breast cancer.
Figures



References
-
- Chang JC, Wooten EC, Tsimelzon A, Hilsenbeck SG, Gutierrez MC, Elledge R, Mohsin S, Osborne CK, Chamness GC, Allred DC, O'Connell P. Gene expression profiling for the prediction of therapeutic response to docetaxel in patients with breast cancer. Lancet. 2003;362:362–369. doi: 10.1016/S0140-6736(03)14023-8. - DOI - PubMed
-
- Ma XJ, Wang Z, Ryan PD, Isakoff SJ, Barmettler A, Fuller A, Muir B, Mohapatra G, Salunga R, Tuggle JT, Tran Y, Tran D, Tassin A, Amon P, Wang W, Wang W, Enright E, Stecker K, Estepa-Sabal E, Smith B, Younger J, Balis U, Michaelson J, Bhan A, Habin K, Baer TM, Brugge J, Haber DA, Erlander MG, Sgroi DC. A two-gene expression ratio predicts clinical outcome in breast cancer patients treated with tamoxifen. Cancer Cell. 2004;5:607–616. doi: 10.1016/j.ccr.2004.05.015. - DOI - PubMed
-
- Sotiriou C, Neo SY, McShane LM, Korn EL, Long PM, Jazaeri A, Martiat P, Fox SB, Harris AL, Liu ET. Breast cancer classification and prognosis based on gene expression profiles from a population-based study. Proc Natl Acad Sci U S A. 2003;100:10393–10398. doi: 10.1073/pnas.1732912100. - DOI - PMC - PubMed
-
- Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, Hastie T, Eisen MB, Van de RM, Jeffrey SS, Thorsen T, Quist H, Matese JC, Brown PO, Botstein D, Eystein LP, Borresen-Dale AL. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci U S A. 2001;98:10869–10874. doi: 10.1073/pnas.191367098. - DOI - PMC - PubMed
-
- van't Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M, Peterse HL, van der KK, Marton MJ, Witteveen AT, Schreiber GJ, Kerkhoven RM, Roberts C, Linsley PS, Bernards R, Friend SH. Gene expression profiling predicts clinical outcome of breast cancer. Nature. 2002;415:530–536. doi: 10.1038/415530a. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

