Yersiniabactin production by Pseudomonas syringae and Escherichia coli, and description of a second yersiniabactin locus evolutionary group
- PMID: 16751485
- PMCID: PMC1489633
- DOI: 10.1128/AEM.00119-06
Yersiniabactin production by Pseudomonas syringae and Escherichia coli, and description of a second yersiniabactin locus evolutionary group
Abstract
The siderophore and virulence factor yersiniabactin is produced by Pseudomonas syringae. Yersiniabactin was originally detected by high-pressure liquid chromatography (HPLC); commonly used PCR tests proved ineffective. Yersiniabactin production in P. syringae correlated with the possession of irp1 located in a predicted yersiniabactin locus. Three similarly divergent yersiniabactin locus groups were determined: the Yersinia pestis group, the P. syringae group, and the Photorhabdus luminescens group; yersiniabactin locus organization is similar in P. syringae and P. luminescens. In P. syringae pv. tomato DC3000, the locus has a high GC content (63.4% compared with 58.4% for the chromosome and 60.1% and 60.7% for adjacent regions) but it lacks high-pathogenicity-island features, such as the insertion in a tRNA locus, the integrase, and insertion sequence elements. In P. syringae pv. tomato DC3000 and pv. phaseolicola 1448A, the locus lies between homologues of Psyr_2284 and Psyr_2285 of P. syringae pv. syringae B728a, which lacks the locus. Among tested pseudomonads, a PCR test specific to two yersiniabactin locus groups detected a locus in genospecies 3, 7, and 8 of P. syringae, and DNA hybridization within P. syringae also detected a locus in the pathovars phaseolicola and glycinea. The PCR and HPLC methods enabled analysis of nonpathogenic Escherichia coli. HPLC-proven yersiniabactin-producing E. coli lacked modifications found in irp1 and irp2 in the human pathogen CFT073, and it is not clear whether CFT073 produces yersiniabactin. The study provides clues about the evolution and dispersion of yersiniabactin genes. It describes methods to detect and study yersiniabactin producers, even where genes have evolved.
Figures

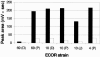

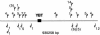
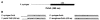
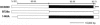
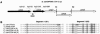
References
-
- Ausubel, F. M., R. Brent, R. E. Kingston, D. D. Moore, J. G. Seidlan, J. A. Smith, and K. Struhl. 2000. Current protocols in molecular biology. John Wiley & Sons, Inc., New York, N.Y.
-
- Bach, S., A. de Almeida, and E. Carniel. 2000. The Yersinia high-pathogenicity island is present in different members of the family Enterobacteriaceae. FEMS Microbiol. Lett. 183:289-294. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

