Automated interpretation of subcellular patterns in fluorescence microscope images for location proteomics
- PMID: 16752421
- PMCID: PMC2901544
- DOI: 10.1002/cyto.a.20280
Automated interpretation of subcellular patterns in fluorescence microscope images for location proteomics
Abstract
Proteomics, the large scale identification and characterization of many or all proteins expressed in a given cell type, has become a major area of biological research. In addition to information on protein sequence, structure and expression levels, knowledge of a protein's subcellular location is essential to a complete understanding of its functions. Currently, subcellular location patterns are routinely determined by visual inspection of fluorescence microscope images. We review here research aimed at creating systems for automated, systematic determination of location. These employ numerical feature extraction from images, feature reduction to identify the most useful features, and various supervised learning (classification) and unsupervised learning (clustering) methods. These methods have been shown to perform significantly better than human interpretation of the same images. When coupled with technologies for tagging large numbers of proteins and high-throughput microscope systems, the computational methods reviewed here enable the new subfield of location proteomics. This subfield will make critical contributions in two related areas. First, it will provide structured, high-resolution information on location to enable Systems Biology efforts to simulate cell behavior from the gene level on up. Second, it will provide tools for Cytomics projects aimed at characterizing the behaviors of all cell types before, during, and after the onset of various diseases.
Copyright 2006 International Society for Analytical Cytology.
Figures
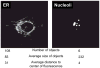



References
-
- Chen X, Velliste M, Weinstein S, Jarvik JW, Murphy RF. Location proteomics - Building subcellular location trees from high resolution 3D fluorescence microscope images of randomly-tagged proteins. Proc SPIE. 2003;4962:298–306.
-
- Boland MV, Markey MK, Murphy RF. Automated recognition of patterns characteristic of subcellular structures in fluorescence microscopy images. Cytometry. 1998;33(3):366–375. - PubMed
-
- Boland MV, Murphy RF. A Neural Network Classifier Capable of Recognizing the Patterns of all Major Subcellular Structures in Fluorescence Microscope Images of HeLa Cells. Bioinformatics. 2001;17(12):1213–1223. - PubMed
-
- Danckaert A, Gonzalez-Couto E, Bollondi L, Thompson N, Hayes B. Automated Recognition of Intracellular Organelles in Confocal Microscope Images. Traffic. 2002;3(1):66–73. - PubMed
-
- Murphy RF, Velliste M, Porreca G. Robust Numerical Features for Description and Classification of Subcellular Location Patterns in Fluorescence Microscope Images. J VLSI Sig Proc. 2003;35(3):311–321.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

