Identification and analysis of DNA-binding transcription factors in Bacillus subtilis and other Firmicutes--a genomic approach
- PMID: 16772031
- PMCID: PMC1524751
- DOI: 10.1186/1471-2164-7-147
Identification and analysis of DNA-binding transcription factors in Bacillus subtilis and other Firmicutes--a genomic approach
Abstract
Background: Bacillus subtilis is one of the best-characterized organisms in Gram-positive bacteria. It represents a paradigm of gene regulation in bacteria due its complex life style (which could involve a transition between stages as diverse as vegetative cell and spore formation). In order to gain insight into the organization and evolution of the B. subtilis regulatory network and to provide an alternative framework for further studies in bacteria, we identified and analyzed its repertoire of DNA-binding transcription factors in terms of their abundance, family distribution and regulated genes.
Results: A collection of 237 DNA-binding Transcription Factors (TFs) was identified in B. subtilis, half of them with experimental evidence. 59% of them were predicted to be repressors, 17% activators, 17% were putatively identified as dual regulatory proteins and the remaining 6.3% could not be associated with a regulatory role. From this collection 56 TFs were found to be autoregulated, most of them negatively, though a significant proportion of positive feedback circuits were also identified. TFs were clustered into 51 regulatory protein families and then traced on 58 genomes from Firmicutes to detect their presence. From this analysis three families were found conserved in all the Firmicutes; fifteen families were distributed in all Firmicutes except in the phyla Mollicutes; two were constrained to Bacillales and finally two families were found to be specific to B. subtilis, due to their specie specific distribution. Repression seems to be the most common regulatory mechanism in Firmicutes due to the high proportion of repressors in the detected collection in these genomes. In addition, six global regulators were defined in B. subtilis based on the number and function of their regulated genes.
Conclusion: In this work we identified and described the characteristics associated to the repertoire of DNA-binding TFs in B. subtilis. We also quantified their abundance, family distribution, and regulatory roles in the context of Firmicutes. This work should not only contribute to our understanding of the regulation of gene expression in bacteria from the perspective of B. subtilis but also provide us the basis for comprehensive modeling of transcriptional regulatory networks in Firmicutes.
Figures

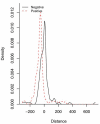


References
-
- Kunst F, Ogasawara N, Moszer I, Albertini AM, Alloni G, Azevedo V, Bertero MG, Bessieres P, Bolotin A, Borchert S, Borriss R, Boursier L, Brans A, Braun M, Brignell SC, Bron S, Brouillet S, Bruschi CV, Caldwell B, Capuano V, Carter NM, Choi SK, Codani JJ, Connerton IF, Danchin A, et al. The complete genome sequence of the gram-positive bacterium Bacillus subtilis. Nature. 1997;390:249–256. doi: 10.1038/36786. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

