Genome Annotation Transfer Utility (GATU): rapid annotation of viral genomes using a closely related reference genome
- PMID: 16772042
- PMCID: PMC1534038
- DOI: 10.1186/1471-2164-7-150
Genome Annotation Transfer Utility (GATU): rapid annotation of viral genomes using a closely related reference genome
Abstract
Background: Since DNA sequencing has become easier and cheaper, an increasing number of closely related viral genomes have been sequenced. However, many of these have been deposited in GenBank without annotations, severely limiting their value to researchers. While maintaining comprehensive genomic databases for a set of virus families at the Viral Bioinformatics Resource Center http://www.biovirus.org and Viral Bioinformatics - Canada http://www.virology.ca, we found that researchers were unnecessarily spending time annotating viral genomes that were close relatives of already annotated viruses. We have therefore designed and implemented a novel tool, Genome Annotation Transfer Utility (GATU), to transfer annotations from a previously annotated reference genome to a new target genome, thereby greatly reducing this laborious task.
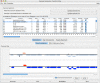
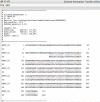
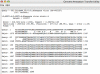
Results: GATU transfers annotations from a reference genome to a closely related target genome, while still giving the user final control over which annotations should be included. GATU also detects open reading frames present in the target but not the reference genome and provides the user with a variety of bioinformatics tools to quickly determine if these ORFs should also be included in the annotation. After this process is complete, GATU saves the newly annotated genome as a GenBank, EMBL or XML-format file. The software is coded in Java and runs on a variety of computer platforms. Its user-friendly Graphical User Interface is specifically designed for users trained in the biological sciences.
Conclusion: GATU greatly simplifies the initial stages of genome annotation by using a closely related genome as a reference. It is not intended to be a gene prediction tool or a "complete" annotation system, but we have found that it significantly reduces the time required for annotation of genes and mature peptides as well as helping to standardize gene names between related organisms by transferring reference genome annotations to the target genome. The program is freely available under the General Public License and can be accessed along with documentation and tutorial from http://www.virology.ca/gatu.
Figures



References
-
- Viral Bioinformatics Resource Center http://www.virology.ca
Publication types
MeSH terms
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

