The conserved AAUAAA hexamer of the poly(A) signal can act alone to trigger a stable decrease in RNA polymerase II transcription velocity
- PMID: 16775304
- PMCID: PMC1524889
- DOI: 10.1261/rna.103206
The conserved AAUAAA hexamer of the poly(A) signal can act alone to trigger a stable decrease in RNA polymerase II transcription velocity
Abstract
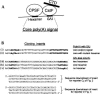
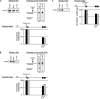
In vivo the poly(A) signal not only directs 3'-end processing but also controls the rate and extent of transcription. Thus, upon crossing the poly(A) signal RNA polymerase II first pauses and then terminates. We show that the G/U-rich region of the poly(A) signal, although required for termination in vivo, is not required for poly(A)-dependent pausing either in vivo or in vitro. Consistent with this, neither CstF, which recognizes the G/U-rich element, nor the polymerase CTD, which binds CstF, is required for pausing. The only part of the poly(A) signal required to direct the polymerase to pause is the AAUAAA hexamer. The effect of the hexamer on the polymerase is long lasting--in many situations polymerases over 1 kb downstream of the hexamer continue to exhibit delayed progress down the template in vivo. The hexamer is the first part of the poly(A) signal to emerge from the polymerase and may play a role independent of the rest of the poly(A) signal in paving the way for subsequent events such as 3'-end processing and termination of transcription.
Figures



Similar articles
-
The poly(A)-dependent transcriptional pause is mediated by CPSF acting on the body of the polymerase.Nat Struct Mol Biol. 2007 Jul;14(7):662-9. doi: 10.1038/nsmb1253. Epub 2007 Jun 17. Nat Struct Mol Biol. 2007. PMID: 17572685
-
3' RNA processing efficiency plays a primary role in generating termination-competent RNA polymerase II elongation complexes.Mol Cell Biol. 1993 Jun;13(6):3472-80. doi: 10.1128/mcb.13.6.3472-3480.1993. Mol Cell Biol. 1993. PMID: 7684499 Free PMC article.
-
A functional mRNA polyadenylation signal is required for transcription termination by RNA polymerase II.Genes Dev. 1988 Apr;2(4):440-52. doi: 10.1101/gad.2.4.440. Genes Dev. 1988. PMID: 2836265
-
Transcription termination and 3' processing: the end is in site!Cell. 1985 Jun;41(2):349-59. doi: 10.1016/s0092-8674(85)80007-6. Cell. 1985. PMID: 2580642 Review. No abstract available.
-
Transcriptional termination in mammals: Stopping the RNA polymerase II juggernaut.Science. 2016 Jun 10;352(6291):aad9926. doi: 10.1126/science.aad9926. Science. 2016. PMID: 27284201 Free PMC article. Review.
Cited by
-
Temporal Dissection of Rate Limiting Transcriptional Events Using Pol II ChIP and RNA Analysis of Adrenergic Stress Gene Activation.PLoS One. 2015 Aug 5;10(8):e0134442. doi: 10.1371/journal.pone.0134442. eCollection 2015. PLoS One. 2015. PMID: 26244980 Free PMC article.
-
Fail-safe transcription termination: Because one is never enough.RNA Biol. 2015;12(9):927-32. doi: 10.1080/15476286.2015.1073433. RNA Biol. 2015. PMID: 26273910 Free PMC article. Review.
-
RNA polymerase II pauses and associates with pre-mRNA processing factors at both ends of genes.Nat Struct Mol Biol. 2008 Jan;15(1):71-8. doi: 10.1038/nsmb1352. Epub 2007 Dec 23. Nat Struct Mol Biol. 2008. PMID: 18157150 Free PMC article.
-
A genomic analysis of RNA polymerase II modification and chromatin architecture related to 3' end RNA polyadenylation.Genome Res. 2008 Aug;18(8):1224-37. doi: 10.1101/gr.075804.107. Epub 2008 May 16. Genome Res. 2008. PMID: 18487515 Free PMC article.
-
Genome-wide mapping of yeast RNA polymerase II termination.PLoS Genet. 2014 Oct 9;10(10):e1004632. doi: 10.1371/journal.pgen.1004632. eCollection 2014 Oct. PLoS Genet. 2014. PMID: 25299594 Free PMC article.
References
-
- Bird G., Fong N., Gatlin J.C., Farabaugh S., Bentley D.L. Ribozyme cleavage reveals connections between mRNA release from the site of transcription and Pre-mRNA processing. Mol. Cell. 2005;20:747–758. - PubMed
-
- Calvo O., Manley J.L. Strange bedfellows: Polyadenylation factors at the promoter. Genes & Dev. 2003;17:1321–1327. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
