Quantitative trait loci for locomotor behavior in Drosophila melanogaster
- PMID: 16783013
- PMCID: PMC1569784
- DOI: 10.1534/genetics.106.058099
Quantitative trait loci for locomotor behavior in Drosophila melanogaster
Abstract
Locomotion is an integral component of most animal behaviors and many human diseases and disorders are associated with locomotor deficits, but little is known about the genetic basis of natural variation in locomotor behavior. Locomotion is a complex trait, with variation attributable to the joint segregation of multiple interacting quantitative trait loci (QTL), with effects that are sensitive to the environment. We assessed variation in a component of locomotor behavior (locomotor reactivity) in a population of 98 recombinant inbred lines of Drosophila melanogaster and mapped four QTL affecting locomotor reactivity by linkage to polymorphic roo transposable element insertion sites. We used complementation tests of deficiencies to fine map these QTL to 12 chromosomal regions and complementation tests of mutations to identify 13 positional candidate genes affecting locomotor reactivity, including Dopa decarboxylase (Ddc), which catalyzes the final step in the synthesis of serotonin and dopamine. Linkage disequilibrium mapping in a population of 164 second chromosome substitution lines derived from a single natural population showed that polymorphisms at Ddc were associated with naturally occurring genetic variation in locomotor behavior. These data implicate variation in the synthesis of bioamines as a factor contributing to natural variation in locomotor reactivity.
Figures






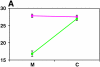













References
-
- Adams, M. D., S. E. Celniker, R. A. Holt, C. A. Evans, J. D. Gocayne et al., 2000. The genome sequence of Drosophila melanogaster. Science 287: 2185–2195. - PubMed
-
- American Psychiatric Association, 1994. Diagnostic and Statistical Manual of Mental Disorders, Ed. 4. American Psychiatric Press, Arlington, VA.
-
- Baker, B. S., B. J. Taylor and J. C. Hall, 2001. Are complex behaviors specified by dedicated regulatory genes? Reasoning from Drosophila. Cell 105: 13–24. - PubMed
-
- Basten, C. J., B. S. Weir and Z. B. Zeng, 1994. Zmap—a QTL cartographer, pp. 65–66 in Proceedings of the 5th World Congress on Genetics Applied to Livestock Production: Computing Strategies and Software, edited by C. Smith, J. S. Gavora, B. Benkel, J. Chesnias and W. Fairfull. Organizing Committee, 5th World Congress on Applied Genetics Applied to Livestock Production, Guelph, Ontario, Canada.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

