Sorting of cells of the same size, shape, and cell cycle stage for a single cell level assay without staining
- PMID: 16790072
- PMCID: PMC1523334
- DOI: 10.1186/1471-2121-7-25
Sorting of cells of the same size, shape, and cell cycle stage for a single cell level assay without staining
Abstract
Background: Single-cell level studies are being used increasingly to measure cell properties not directly observable in a cell population. High-performance data acquisition systems for such studies have, by necessity, developed in synchrony. However, improvements in sample purification techniques are also required to reveal new phenomena. Here we assessed a cell sorter as a sample-pretreatment tool for a single-cell level assay. A cell sorter is routinely used for selecting one type of cells from a heterogeneous mixture of cells using specific fluorescence labels. In this case, we wanted to select cells of exactly the same size, shape, and cell-cycle stage from a population, without using a specific fluorescence label.
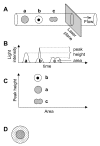
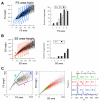
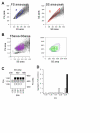
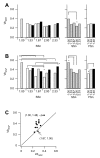
Results: We used four light scatter parameters: the peak height and area of the forward scatter (FSheight and FSarea) and side scatter (SSheight and SSarea). The rat pheochromocytoma PC12 cell line, a neuronal cell line, was used for all experiments. The living cells concentrated in the high FSarea and middle SSheight/SSarea fractions. Single cells without cell clumps were concentrated in the low SS and middle FS fractions, and in the higher FSheight/FSarea and SSheight/SSarea fractions. The cell populations from these viable, single-cell-rich fractions were divided into twelve subfractions based on their FSarea-SSarea profiles, for more detailed analysis. We found that SSarea was proportional to the cell volume and the FSarea correlated with cell roundness and elongation, as well as with the level of DNA in the cell. To test the method and to characterize the basic properties of the isolated single cells, sorted cells were cultured in separate wells. The cells in all subfractions survived, proliferated and differentiated normally, suggesting that there was no serious damage. The smallest, roundest, and smoothest cells had the highest viability. There was no correlation between proliferation and differentiation. NGF increases cell viability but decreases the proliferative ability of the PC12 cells.
Conclusion: We demonstrated a pretreatment method to collect well-characterized, viable, single cells without using fluorescent labels and without significant damage to the cells. This method is quantitative, rapid, single-step, and yields cells of high purity, making it applicable for a variety of single-cell level analyses.
Figures


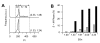
References
-
- Salzman GC, Crowell JM, Martin JC, Trujillo TT, Romero A, Mullaney PF, LaBauve PM. Cell classification by laser light scattering: identification and separation of unstained leukocytes. Acta Cytol. 1975;19:374–377. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

