Hosting the severe acute respiratory syndrome coronavirus: specific cell factors required for infection
- PMID: 16803585
- PMCID: PMC7162409
- DOI: 10.1111/j.1462-5822.2006.00744.x
Hosting the severe acute respiratory syndrome coronavirus: specific cell factors required for infection
Abstract
As with all viruses, the severe acute respiratory syndrome coronavirus (SARS-CoV) utilizes specific host cell factors during its infection cycle. Some of these factors have been identified and are now increasingly scrutinized as targets to intervene with infection. In this brief review, we describe the current understanding of how the SARS-CoV is able to use the cellular machinery for its replication.
Figures
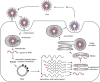
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

