Role of p38 in replication of Trypanosoma brucei kinetoplast DNA
- PMID: 16809774
- PMCID: PMC1592711
- DOI: 10.1128/MCB.00369-06
Role of p38 in replication of Trypanosoma brucei kinetoplast DNA
Abstract
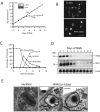
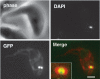
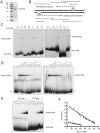
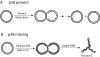
Trypanosomes have an unusual mitochondrial genome, called kinetoplast DNA, that is a giant network containing thousands of interlocked minicircles. During kinetoplast DNA synthesis, minicircles are released from the network for replication as theta-structures, and then the free minicircle progeny reattach to the network. We report that a mitochondrial protein, which we term p38, functions in kinetoplast DNA replication. RNA interference (RNAi) of p38 resulted in loss of kinetoplast DNA and accumulation of a novel free minicircle species named fraction S. Fraction S minicircles are so underwound that on isolation they become highly negatively supertwisted and develop a region of Z-DNA. p38 binds to minicircle sequences within the replication origin. We conclude that cells with RNAi-induced loss of p38 cannot initiate minicircle replication, although they can extensively unwind free minicircles.
Figures



Similar articles
-
A second mitochondrial DNA primase is essential for cell growth and kinetoplast minicircle DNA replication in Trypanosoma brucei.Eukaryot Cell. 2011 Mar;10(3):445-54. doi: 10.1128/EC.00308-10. Epub 2011 Jan 21. Eukaryot Cell. 2011. PMID: 21257796 Free PMC article.
-
TbPIF1, a Trypanosoma brucei mitochondrial DNA helicase, is essential for kinetoplast minicircle replication.J Biol Chem. 2010 Mar 5;285(10):7056-66. doi: 10.1074/jbc.M109.084038. Epub 2009 Dec 30. J Biol Chem. 2010. PMID: 20042610 Free PMC article.
-
Stem-loop silencing reveals that a third mitochondrial DNA polymerase, POLID, is required for kinetoplast DNA replication in trypanosomes.Eukaryot Cell. 2008 Dec;7(12):2141-6. doi: 10.1128/EC.00199-08. Epub 2008 Oct 10. Eukaryot Cell. 2008. PMID: 18849470 Free PMC article.
-
Fellowship of the rings: the replication of kinetoplast DNA.Trends Parasitol. 2005 Aug;21(8):363-9. doi: 10.1016/j.pt.2005.06.008. Trends Parasitol. 2005. PMID: 15967722 Review.
-
Closing the gaps in kinetoplast DNA network replication.Proc Natl Acad Sci U S A. 2004 Mar 30;101(13):4333-4. doi: 10.1073/pnas.0401400101. Epub 2004 Mar 22. Proc Natl Acad Sci U S A. 2004. PMID: 15070715 Free PMC article. Review. No abstract available.
Cited by
-
The Trypanosoma cruzi nucleic acid binding protein Tc38 presents changes in the intramitochondrial distribution during the cell cycle.BMC Microbiol. 2009 Feb 11;9:34. doi: 10.1186/1471-2180-9-34. BMC Microbiol. 2009. PMID: 19210781 Free PMC article.
-
A second mitochondrial DNA primase is essential for cell growth and kinetoplast minicircle DNA replication in Trypanosoma brucei.Eukaryot Cell. 2011 Mar;10(3):445-54. doi: 10.1128/EC.00308-10. Epub 2011 Jan 21. Eukaryot Cell. 2011. PMID: 21257796 Free PMC article.
-
Characterization of the novel mitochondrial genome replication factor MiRF172 in Trypanosoma brucei.J Cell Sci. 2018 Apr 25;131(8):jcs211730. doi: 10.1242/jcs.211730. J Cell Sci. 2018. PMID: 29626111 Free PMC article.
-
U-insertion/deletion RNA editing multiprotein complexes and mitochondrial ribosomes in Leishmania tarentolae are located in antipodal nodes adjacent to the kinetoplast DNA.Mitochondrion. 2015 Nov;25:76-86. doi: 10.1016/j.mito.2015.10.006. Epub 2015 Oct 14. Mitochondrion. 2015. PMID: 26462764 Free PMC article.
-
Mitochondrial origin-binding protein UMSBP mediates DNA replication and segregation in trypanosomes.Proc Natl Acad Sci U S A. 2007 Dec 4;104(49):19250-5. doi: 10.1073/pnas.0706858104. Epub 2007 Nov 28. Proc Natl Acad Sci U S A. 2007. PMID: 18048338 Free PMC article.
References
-
- Brun, R., and M. Schonenberger. 1979. Cultivation and in vitro cloning of procyclic culture forms of Trypanosoma brucei in a semi-defined medium. Acta Trop. 36:289-292. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
