Functional and physical interaction of yeast Mgs1 with PCNA: impact on RAD6-dependent DNA damage tolerance
- PMID: 16809783
- PMCID: PMC1592726
- DOI: 10.1128/MCB.00307-06
Functional and physical interaction of yeast Mgs1 with PCNA: impact on RAD6-dependent DNA damage tolerance
Abstract
Proliferating cell nuclear antigen (PCNA), a sliding clamp required for processive DNA synthesis, provides attachment sites for various other proteins that function in DNA replication, DNA repair, cell cycle progression and chromatin assembly. It has been shown that differential posttranslational modifications of PCNA by ubiquitin or SUMO play a pivotal role in controlling the choice of pathway for rescuing stalled replication forks. Here, we explored the roles of Mgs1 and PCNA in replication fork rescue. We provide evidence that Mgs1 physically associates with PCNA and that Mgs1 helps suppress the RAD6 DNA damage tolerance pathway in the absence of exogenous DNA damage. We also show that PCNA sumoylation inhibits the growth of mgs1 rad18 double mutants, in which PCNA sumoylation and the Srs2 DNA helicase coordinately prevent RAD52-dependent homologous recombination. The proposed roles for Mgs1, Srs2, and modified PCNA during replication arrest highlight the importance of modulating the RAD6 and RAD52 pathways to avoid genome instability.
Figures


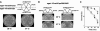


Similar articles
-
Saccharomyces cerevisiae MGS1 is essential in strains deficient in the RAD6-dependent DNA damage tolerance pathway.EMBO J. 2002 Apr 15;21(8):2019-29. doi: 10.1093/emboj/21.8.2019. EMBO J. 2002. PMID: 11953321 Free PMC article.
-
SUMO-modified PCNA recruits Srs2 to prevent recombination during S phase.Nature. 2005 Jul 21;436(7049):428-33. doi: 10.1038/nature03665. Epub 2005 Jun 1. Nature. 2005. PMID: 15931174
-
Error-free DNA-damage tolerance in Saccharomyces cerevisiae.Mutat Res Rev Mutat Res. 2015 Apr-Jun;764:43-50. doi: 10.1016/j.mrrev.2015.02.001. Epub 2015 Feb 16. Mutat Res Rev Mutat Res. 2015. PMID: 26041265 Review.
-
Opposing effects of ubiquitin conjugation and SUMO modification of PCNA on replicational bypass of DNA lesions in Saccharomyces cerevisiae.Mol Cell Biol. 2004 May;24(10):4267-74. doi: 10.1128/MCB.24.10.4267-4274.2004. Mol Cell Biol. 2004. PMID: 15121847 Free PMC article.
-
How yeast cells deal with stalled replication forks.Curr Genet. 2020 Oct;66(5):911-915. doi: 10.1007/s00294-020-01082-y. Epub 2020 May 11. Curr Genet. 2020. PMID: 32394094 Review.
Cited by
-
Post-translational modifications of proliferating cell nuclear antigen: A key signal integrator for DNA damage response (Review).Oncol Lett. 2014 May;7(5):1363-1369. doi: 10.3892/ol.2014.1943. Epub 2014 Mar 5. Oncol Lett. 2014. PMID: 24765138 Free PMC article.
-
Prevention of unwanted recombination at damaged replication forks.Curr Genet. 2020 Dec;66(6):1045-1051. doi: 10.1007/s00294-020-01095-7. Epub 2020 Jul 15. Curr Genet. 2020. PMID: 32671464 Free PMC article. Review.
-
Bacillus subtilis RarA modulates replication restart.Nucleic Acids Res. 2018 Aug 21;46(14):7206-7220. doi: 10.1093/nar/gky541. Nucleic Acids Res. 2018. PMID: 29947798 Free PMC article.
-
The AAA+ superfamily of functionally diverse proteins.Genome Biol. 2008 Apr 30;9(4):216. doi: 10.1186/gb-2008-9-4-216. Genome Biol. 2008. PMID: 18466635 Free PMC article. Review.
-
A novel variant of DNA polymerase ζ, Rev3ΔC, highlights differential regulation of Pol32 as a subunit of polymerase δ versus ζ in Saccharomyces cerevisiae.DNA Repair (Amst). 2014 Dec;24:138-149. doi: 10.1016/j.dnarep.2014.04.013. Epub 2014 May 10. DNA Repair (Amst). 2014. PMID: 24819597 Free PMC article.
References
-
- Aroya, S. B., and M. Kupiec. 2005. The Elg1 replication factor C-like complex: a novel guardian of genome stability. DNA Repair 4:409-417. - PubMed
-
- Bailly, V., J. Lamb, P. Sung, S. Prakash, and L. Prakash. 1994. Specific complex formation between yeast RAD6 and RAD18 proteins: a potential mechanism for targeting RAD6 ubiquitin-conjugating activity to DNA damage sites. Genes Dev. 8:811-820. - PubMed
-
- Bailly, V., S. Lauder, S. Prakash, and L. Prakash. 1997. Yeast DNA repair proteins Rad6 and Rad18 form a heterodimer that has ubiquitin conjugating, DNA binding, and ATP hydrolytic activities. J. Biol. Chem. 272:23360-23365. - PubMed
-
- Barbour, L., and W. Xiao. 2003. Regulation of alternative replication bypass pathways at stalled replication forks and its effects on genome stability: a yeast model. Mutat. Res. 532:137-155. - PubMed
-
- Branzei, D., M. Seki, F. Onoda, and T. Enomoto. 2002. The product of Saccharomyces cerevisiae WHIP/MGS1, a gene related to replication factor C genes, interacts functionally with DNA polymerase delta. Mol. Genet. Genomics 268:371-386. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
