Phylogenomic and biochemical characterization of three Legionella pneumophila polypeptide deformylases
- PMID: 16816197
- PMCID: PMC1539947
- DOI: 10.1128/JB.00866-05
Phylogenomic and biochemical characterization of three Legionella pneumophila polypeptide deformylases
Abstract
Legionella pneumophila is a gram-negative facultative intracellular human pathogen that can cause fatal Legionnaires' disease. Polypeptide deformylase (PDF) is a novel broad-spectrum antibacterial target, and reports of inhibitors of PDF with potent activities against L. pneumophila have been published previously. Here, we report the identification of not one but three putative pdf genes, pdfA, pdfB, and pdfC, in the complete genome sequences of three strains of L. pneumophila. Phylogenetic analysis showed that L. pneumophila PdfA is most closely related to the commonly known gamma-proteobacterial PDFs encoded by the gene def. PdfB and PdfC are more divergent and do not cluster with any specific bacterial or eukaryotic PDF. All three putative pdf genes from L. pneumophila strain Philadelphia 1 have been cloned, and their encoded products have been overexpressed in Escherichia coli and purified. Enzymatic characterization shows that the purified PDFs with Ni2+ substituted are catalytically active and able to remove the N-formyl group from several synthetic polypeptides, although they appear to have different substrate specificities. Surprisingly, while PdfA and PdfB with Zn2+ substituted are much less active than the Ni2+ forms of each enzyme, PdfC with Zn2+ substituted was as active as the Ni2+ form for the fMA substrate and exhibited substrate specificity different from that of Ni2+ PdfC. Furthermore, the catalytic activities of these enzymes are potently inhibited by a known small-molecule PDF inhibitor, BB-3497, which also inhibits the extracellular growth of L. pneumophila. These results indicate that even though L. pneumophila has three PDFs, they can be effectively inhibited by PDF inhibitors which can, therefore, have potent anti-L. pneumophila activity.
Figures


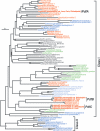
 ). The scale bar represents 0.1 expected amino acid residue substitution per site.
). The scale bar represents 0.1 expected amino acid residue substitution per site.Similar articles
-
The origins of eukaryotic-like proteins in Legionella pneumophila.Int J Med Microbiol. 2010 Nov;300(7):470-81. doi: 10.1016/j.ijmm.2010.04.016. Epub 2010 May 26. Int J Med Microbiol. 2010. PMID: 20537944
-
Characterization of an acetyltransferase that detoxifies aromatic chemicals in Legionella pneumophila.Biochem J. 2012 Jul 15;445(2):219-28. doi: 10.1042/BJ20120528. Biochem J. 2012. PMID: 22545684
-
Purification and characterization of a UDP-glucosyltransferase produced by Legionella pneumophila.Infect Immun. 2003 Jan;71(1):181-6. doi: 10.1128/IAI.71.1.181-186.2003. Infect Immun. 2003. PMID: 12496164 Free PMC article.
-
Legionella pneumophila: population genetics, phylogeny and genomics.Infect Genet Evol. 2009 Sep;9(5):727-39. doi: 10.1016/j.meegid.2009.05.004. Epub 2009 May 18. Infect Genet Evol. 2009. PMID: 19450709 Review.
-
Characterisation of Legionella pneumophila phospholipases and their impact on host cells.Eur J Cell Biol. 2011 Nov;90(11):903-12. doi: 10.1016/j.ejcb.2010.12.003. Epub 2011 Feb 20. Eur J Cell Biol. 2011. PMID: 21342713 Review.
Cited by
-
Identification of crucial amino acids of bacterial Peptide deformylases affecting enzymatic activity in response to oxidative stress.J Bacteriol. 2014 Jan;196(1):90-9. doi: 10.1128/JB.00916-13. Epub 2013 Oct 18. J Bacteriol. 2014. PMID: 24142250 Free PMC article.
-
Structures of Legionella pneumophila serogroup 1 peptide deformylase bound to nickel(II) and actinonin.Acta Crystallogr F Struct Biol Commun. 2025 Apr 1;81(Pt 4):163-170. doi: 10.1107/S2053230X25001876. Epub 2025 Mar 17. Acta Crystallogr F Struct Biol Commun. 2025. PMID: 40091854
-
In vitro and intracellular activities of peptide deformylase inhibitor GSK1322322 against Legionella pneumophila isolates.Antimicrob Agents Chemother. 2015 Jan;59(1):707-10. doi: 10.1128/AAC.04006-14. Epub 2014 Oct 27. Antimicrob Agents Chemother. 2015. PMID: 25348534 Free PMC article.
-
Ligand and Structure-Based Approaches for the Identification of Peptide Deformylase Inhibitors as Antibacterial Drugs.Int J Mol Sci. 2016 Jul 15;17(7):1141. doi: 10.3390/ijms17071141. Int J Mol Sci. 2016. PMID: 27428963 Free PMC article.
-
The C-terminal residue of phage Vp16 PDF, the smallest peptide deformylase, acts as an offset element locking the active conformation.Sci Rep. 2017 Sep 8;7(1):11041. doi: 10.1038/s41598-017-11329-3. Sci Rep. 2017. PMID: 28887476 Free PMC article.
References
-
- Boularot, A., C. Giglione, I. Artaud, and T. Meinnel. 2004. Structure-activity relationship analysis and therapeutic potential of peptide deformylase inhibitors. Curr. Opin. Investig. Drugs 5:809-822. - PubMed
-
- Brenner, D. J., A. Steigerwalt, J. E. Weaver, J. E. McDade, J. C. Feeley, and M. Mandel. 1978. Classification of the Legionnaires' disease bacterium: an interim report. Curr. Microbiol. 1:71-75.
-
- Cazalet, C., C. Rusniok, H. Bruggemann, N. Zidane, A. Magnier, L. Ma, M. Tichit, S. Jarraud, C. Bouchier, F. Vandenesch, F. Kunst, J. Etienne, P. Glaser, and C. Buchrieser. 2004. Evidence in the Legionella pneumophila genome for exploitation of host cell functions and high genome plasticity. Nat. Genet. 36:1165-1173. - PubMed
-
- Chan, P. F., K. M. O'Dwyer, L. M. Palmer, J. D. Ambrad, K. A. Ingraham, C. So, M. A. Lonetto, S. Biswas, M. Rosenberg, D. J. Holmes, and M. Zalacain. 2003. Characterization of a novel fucose-regulated promoter (PfcsK) suitable for gene essentiality and antibacterial mode-of-action studies in Streptococcus pneumoniae. J. Bacteriol. 185:2051-2058. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

