Alanine dehydrogenase activity is required for adequate progression of phycobilisome degradation during nitrogen starvation in Synechococcus elongatus PCC 7942
- PMID: 16816198
- PMCID: PMC1539948
- DOI: 10.1128/JB.00209-06
Alanine dehydrogenase activity is required for adequate progression of phycobilisome degradation during nitrogen starvation in Synechococcus elongatus PCC 7942
Abstract
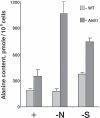
Degradation of the cyanobacterial light-harvesting antenna, the phycobilisome, is a general acclimation response that is observed under various stress conditions. In this study we identified a novel mutant of Synechococcus elongatus PCC 7942 that exhibits impaired phycobilisome degradation specifically during nitrogen starvation, unlike previously described mutants, which exhibit aberrant degradation under nitrogen, sulfur, and phosphorus starvation conditions. The phenotype of the new mutant, AldOmega, results from inactivation of ald (encoding alanine dehydrogenase). AldOmega is deficient in transcription induction of a number of genes during nitrogen starvation. These genes include the "general nutrient stress-related" genes, nblA and nblC, the products of which are essential for phycobilisome degradation. Furthermore, transcripts of several specific nitrogen-responsive genes accumulate at lower levels in AldOmega than in the wild-type strain. In contrast, ald inactivation did not decrease the accumulation of transcripts during sulfur starvation. Transcription of ald is induced upon nitrogen starvation, which is consistent with the ability of wild-type cells to maintain a low cellular content of alanine under these conditions. Unlike wild-type cells, AldOmega accumulates alanine upon nitrogen starvation. Our analyses suggest that alanine dehydrogenase activity is necessary for an adequate cellular response to nitrogen starvation. Decomposition of alanine may be required to provide a sufficient amount of ammonia. Furthermore, the accumulated alanine, or a related metabolite, may interfere with the cues that modulate acclimation during nitrogen starvation. Taken together, our results provide novel information regarding cellular responses to nitrogen starvation and suggest that mechanisms related to nitrogen-specific responses are involved in modulation of a general acclimation process.
Figures

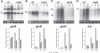

Similar articles
-
NblC, a novel component required for pigment degradation during starvation in Synechococcus PCC 7942.Mol Microbiol. 2005 Nov;58(3):659-68. doi: 10.1111/j.1365-2958.2005.04844.x. Mol Microbiol. 2005. PMID: 16238617
-
Decomposition of cyanobacterial light harvesting complexes: NblA-dependent role of the bilin lyase homolog NblB.Plant J. 2018 Jun;94(5):813-821. doi: 10.1111/tpj.13896. Epub 2018 Apr 17. Plant J. 2018. PMID: 29575252
-
Nitrogen or sulfur starvation differentially affects phycobilisome degradation and expression of the nblA gene in Synechocystis strain PCC 6803.J Bacteriol. 2001 May;183(10):2989-94. doi: 10.1128/JB.183.10.2989-2994.2001. J Bacteriol. 2001. PMID: 11325925 Free PMC article.
-
Alanine dehydrogenases in mycobacteria.J Microbiol. 2019 Feb;57(2):81-92. doi: 10.1007/s12275-019-8543-7. Epub 2019 Jan 31. J Microbiol. 2019. PMID: 30706339 Review.
-
Recovery of Unicellular Cyanobacteria from Nitrogen Chlorosis: A Model for Resuscitation of Dormant Bacteria.Microb Physiol. 2021;31(2):78-87. doi: 10.1159/000515742. Epub 2021 Apr 20. Microb Physiol. 2021. PMID: 33878759 Review.
Cited by
-
Analysis of Fast Fluorescence Kinetics of a Single Cyanobacterium Trapped in an Optical Microcavity.Plants (Basel). 2023 Jan 30;12(3):607. doi: 10.3390/plants12030607. Plants (Basel). 2023. PMID: 36771691 Free PMC article.
-
Structural, functional, and mutational analysis of the NblA protein provides insight into possible modes of interaction with the phycobilisome.J Biol Chem. 2008 Oct 31;283(44):30330-40. doi: 10.1074/jbc.M804241200. Epub 2008 Aug 21. J Biol Chem. 2008. PMID: 18718907 Free PMC article.
-
Phycobilisome breakdown effector NblD is required to maintain the cellular amino acid composition during nitrogen starvation.J Bacteriol. 2021 Jan 1;204(1):JB0015821. doi: 10.1128/JB.00158-21. Epub 2021 Jul 6. J Bacteriol. 2021. PMID: 34228497 Free PMC article.
-
PipY, a Member of the Conserved COG0325 Family of PLP-Binding Proteins, Expands the Cyanobacterial Nitrogen Regulatory Network.Front Microbiol. 2017 Jul 11;8:1244. doi: 10.3389/fmicb.2017.01244. eCollection 2017. Front Microbiol. 2017. PMID: 28744260 Free PMC article.
-
Nitrogen Starvation Acclimation in Synechococcus elongatus: Redox-Control and the Role of Nitrate Reduction as an Electron Sink.Life (Basel). 2015 Mar 13;5(1):888-904. doi: 10.3390/life5010888. Life (Basel). 2015. PMID: 25780959 Free PMC article.
References
-
- Baier, K., H. Lehmann, D. P. Stephan, and W. Lockau. 2004. NblA is essential for phycobilisome degradation in Anabaena sp. strain PCC 7120 but not for development of functional heterocysts. Microbiology 150:2739-2749. - PubMed
-
- Baier, K., S. Nicklisch, C. Grundner, J. Reinecke, and W. Lockau. 2001. Expression of two nblA-homologous genes is required for phycobilisome degradation in nitrogen-starved Synechocystis sp. PCC6803. FEMS Microbiol. Lett. 195:35-39. - PubMed
-
- Bhaya, D., R. Schwarz, and A. R. Grossman. 1997. Molecular responses to environmental stress, p. 397-442. In M. Potts and B. A. Whitton (ed.), Ecology of cyanobacteria: their diversity in time and space. Kluwer Academic Publishers, Dordrecht, The Netherlands.
-
- Bienert, R., K. Baier, R. Volkmer, W. Lockau, and U. Heinemann. 2006. Crystal structure of NblA from Anabaena sp. PCC 7120, a small protein playing a key role in phycobilisome degradation. J. Biol. Chem. 281:5216-5223. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

