Glycogen phosphorylase, the product of the glgP Gene, catalyzes glycogen breakdown by removing glucose units from the nonreducing ends in Escherichia coli
- PMID: 16816199
- PMCID: PMC1539952
- DOI: 10.1128/JB.01566-05
Glycogen phosphorylase, the product of the glgP Gene, catalyzes glycogen breakdown by removing glucose units from the nonreducing ends in Escherichia coli
Abstract
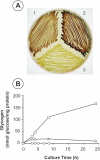
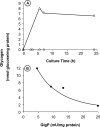
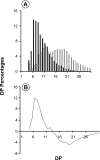
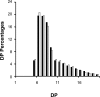
To understand the biological function of bacterial glycogen phosphorylase (GlgP), we have produced and characterized Escherichia coli cells with null or altered glgP expression. glgP deletion mutants (DeltaglgP) totally lacked glycogen phosphorylase activity, indicating that all the enzymatic activity is dependent upon the glgP product. Moderate increases of glycogen phosphorylase activity were accompanied by marked reductions of the intracellular glycogen levels in cells cultured in the presence of glucose. In turn, both glycogen content and rates of glycogen accumulation in DeltaglgP cells were severalfold higher than those of wild-type cells. These defects correlated with the presence of longer external chains in the polysaccharide accumulated by DeltaglgP cells. The overall results thus show that GlgP catalyzes glycogen breakdown and affects glycogen structure by removing glucose units from the polysaccharide outer chains in E. coli.
Figures

References
-
- Ausubel, F. M., R. Brent, R. E. Kingston, D. D. Moore, J. G. Seidman, J. A. Smith, and K. Struhl. 1990. Current protocols in molecular biology. Wiley Interscience, New York, N.Y.
-
- Baker, C. S., I. Morozov, K. Suzuki, T. Romeo, and P. Babitzke. 2002. CsrA regulates glycogen biosynthesis by preventing translation of glgC in Escherichia coli. Mol. Microbiol. 44:1599-1610. - PubMed
-
- Ball, S. G., and M. K. Morell. 2003. From bacterial glycogen to starch: understanding the biogenesis of the plant starch granule. Annu. Rev. Plant Biol. 54:207-233. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

