Antagonism of Chk1 signaling in the G2 DNA damage checkpoint by dominant alleles of Cdr1
- PMID: 16816416
- PMCID: PMC1569782
- DOI: 10.1534/genetics.106.060970
Antagonism of Chk1 signaling in the G2 DNA damage checkpoint by dominant alleles of Cdr1
Abstract
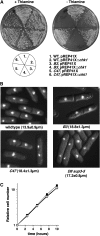
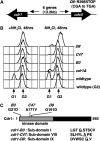
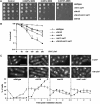
Activation of the Chk1 protein kinase by DNA damage enforces a checkpoint that maintains Cdc2 in its inactive, tyrosine-15 (Y15) phosphorylated state. Chk1 downregulates the Cdc25 phosphatases and concomitantly upregulates the Wee1 kinases that control the phosphorylation of Cdc2. Overproduction of Chk1 causes G(2) arrest/delay independently of DNA damage and upstream checkpoint genes. We utilized this to screen fission yeast for mutations that alter sensitivity to Chk1 signaling. We describe three dominant-negative alleles of cdr1, which render cells supersensitive to Chk1 levels, and suppress the checkpoint defects of chk1Delta cells. Cdr1 encodes a protein kinase previously identified as a negative regulator of Wee1 activity in response to limited nutrition, but Cdr1 has not previously been linked to checkpoint signaling. Overproduction of Cdr1 promotes checkpoint defects and exacerbates the defective response to DNA damage of cells lacking Chk1. We conclude that regulation of Wee1 by Cdr1 and possibly by related kinases is an important antagonist of Chk1 signaling and represents a novel negative regulation of cell cycle arrest promoted by this checkpoint.
Figures



Similar articles
-
Transformation/transcription domain-associated protein (TRRAP)-mediated regulation of Wee1.Genetics. 2010 May;185(1):81-93. doi: 10.1534/genetics.110.114769. Epub 2010 Mar 1. Genetics. 2010. PMID: 20194963 Free PMC article.
-
Chk1 is a wee1 kinase in the G2 DNA damage checkpoint inhibiting cdc2 by Y15 phosphorylation.EMBO J. 1997 Feb 3;16(3):545-54. doi: 10.1093/emboj/16.3.545. EMBO J. 1997. PMID: 9034337 Free PMC article.
-
Two Distinct Cdc2 Pools Regulate Cell Cycle Progression and the DNA Damage Response in the Fission Yeast S.pombe.PLoS One. 2015 Jul 1;10(7):e0130748. doi: 10.1371/journal.pone.0130748. eCollection 2015. PLoS One. 2015. PMID: 26131711 Free PMC article.
-
Establishment of and recovery from damage checkpoint requires sequential interactions of Crb2 with protein kinases Rad3, Chk1, and Cdc2.Cold Spring Harb Symp Quant Biol. 2000;65:443-9. doi: 10.1101/sqb.2000.65.443. Cold Spring Harb Symp Quant Biol. 2000. PMID: 12760060 Review. No abstract available.
-
The G(2) DNA damage checkpoint: could this ancient regulator be the Achilles heel of cancer?Cancer Biol Ther. 2009 Aug;8(15):1433-9. doi: 10.4161/cbt.8.15.9081. Epub 2009 Aug 22. Cancer Biol Ther. 2009. PMID: 19574738 Review.
Cited by
-
Brc1-dependent recovery from replication stress.J Cell Sci. 2012 Jun 1;125(Pt 11):2753-64. doi: 10.1242/jcs.103119. Epub 2012 Feb 24. J Cell Sci. 2012. PMID: 22366461 Free PMC article.
-
Transformation/transcription domain-associated protein (TRRAP)-mediated regulation of Wee1.Genetics. 2010 May;185(1):81-93. doi: 10.1534/genetics.110.114769. Epub 2010 Mar 1. Genetics. 2010. PMID: 20194963 Free PMC article.
-
XPG-related nucleases are hierarchically recruited for double-stranded rDNA break resection.J Biol Chem. 2019 May 10;294(19):7632-7643. doi: 10.1074/jbc.RA118.005415. Epub 2019 Mar 18. J Biol Chem. 2019. PMID: 30885940 Free PMC article.
-
Regulation of chk1.Cell Div. 2009 Apr 29;4:8. doi: 10.1186/1747-1028-4-8. Cell Div. 2009. PMID: 19400965 Free PMC article.
-
Smc5-Smc6-dependent removal of cohesin from mitotic chromosomes.Mol Cell Biol. 2009 Aug;29(16):4363-75. doi: 10.1128/MCB.00377-09. Epub 2009 Jun 15. Mol Cell Biol. 2009. PMID: 19528228 Free PMC article.
References
-
- Barbet, N. C., and A. M. Carr, 1993. Fission yeast wee1 protein kinase is not required for DNA damage-dependent mitotic arrest. Nature 364: 824–827. - PubMed
-
- Barbet, N., W. J. Muriel and A. M. Carr, 1992. Versatile shuttle vectors and genomic libraries for use with Schizosaccharomyces pombe. Gene 114: 59–66. - PubMed
-
- Barr, F. A., H. H. Sillje and E. A. Nigg, 2004. Polo-like kinases and the orchestration of cell division. Nat. Rev. Mol. Cell Biol. 5: 429–440. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

