Gene conversion between direct noncoding repeats promotes genetic and phenotypic diversity at a regulatory locus of Zea mays (L.)
- PMID: 16816430
- PMCID: PMC1602084
- DOI: 10.1534/genetics.105.053942
Gene conversion between direct noncoding repeats promotes genetic and phenotypic diversity at a regulatory locus of Zea mays (L.)
Abstract
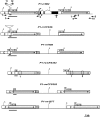
While evolution of coding sequences has been intensively studied, diversification of noncoding regulatory regions remains poorly understood. In this study, we investigated the molecular evolution of an enhancer region located 5 kb upstream of the transcription start site of the maize pericarp color1 (p1) gene. The p1 gene encodes an R2R3 Myb-like transcription factor that regulates the flavonoid biosynthetic pathway in maize floral organs. Distinct p1 alleles exhibit organ-specific expression patterns on kernel pericarp and cob glumes. A cob glume-specific regulatory region has been identified in the distal enhancer. Further characterization of 6 single-copy p1 alleles, including P1-rr (red pericarp/red cob) and P1-rw (red pericarp and white cob), reveals 3 distinct enhancer types. Sequence variations in the enhancer are correlated with the p1 gene expression patterns in cob glume. Structural comparisons and phylogenetic analyses suggest that evolution of the enhancer region is likely driven by gene conversion between long direct noncoding repeats (approximately 6 kb in length). Given that tandem and segmental duplications are common in both animal and plant genomes, our studies suggest that recombination between noncoding duplicated sequences could play an important role in creating genetic and phenotypic variations.
Figures





Similar articles
-
Gene structure induced epigenetic modifications of pericarp color1 alleles of maize result in tissue-specific mosaicism.PLoS One. 2009 Dec 14;4(12):e8231. doi: 10.1371/journal.pone.0008231. PLoS One. 2009. PMID: 20011605 Free PMC article.
-
Comparisons of maize pericarp color1 alleles reveal paralogous gene recombination and an organ-specific enhancer region.Plant Cell. 2005 Mar;17(3):903-14. doi: 10.1105/tpc.104.029660. Epub 2005 Feb 18. Plant Cell. 2005. PMID: 15722466 Free PMC article.
-
Epigenetic modifications of distinct sequences of the p1 regulatory gene specify tissue-specific expression patterns in maize.Genetics. 2007 Mar;175(3):1059-70. doi: 10.1534/genetics.106.066134. Epub 2006 Dec 18. Genetics. 2007. PMID: 17179091 Free PMC article.
-
Progressive loss of DNA methylation releases epigenetic gene silencing from a tandemly repeated maize Myb gene.Genetics. 2009 Jan;181(1):81-91. doi: 10.1534/genetics.108.097170. Epub 2008 Nov 10. Genetics. 2009. PMID: 19001287 Free PMC article.
-
Tissue-specific patterns of a maize Myb transcription factor are epigenetically regulated.Plant J. 2001 Sep;27(5):467-78. doi: 10.1046/j.1365-313x.2001.01124.x. Plant J. 2001. PMID: 11576430
Cited by
-
Gene structure induced epigenetic modifications of pericarp color1 alleles of maize result in tissue-specific mosaicism.PLoS One. 2009 Dec 14;4(12):e8231. doi: 10.1371/journal.pone.0008231. PLoS One. 2009. PMID: 20011605 Free PMC article.
-
Transcribed enhancer sequences are required for maize p1 paramutation.Genetics. 2024 Jan 3;226(1):iyad178. doi: 10.1093/genetics/iyad178. Genetics. 2024. PMID: 38169343 Free PMC article.
-
Heterogeneous tempo and mode of conserved noncoding sequence evolution among four mammalian orders.Genome Biol Evol. 2013;5(12):2330-43. doi: 10.1093/gbe/evt177. Genome Biol Evol. 2013. PMID: 24259317 Free PMC article.
-
Divergence of gene regulation through chromosomal rearrangements.BMC Genomics. 2010 Nov 30;11:678. doi: 10.1186/1471-2164-11-678. BMC Genomics. 2010. PMID: 21118519 Free PMC article.
-
Genetic analysis of pericarp pigmentation variation in Corn Belt dent maize.G3 (Bethesda). 2023 Dec 29;14(1):jkad256. doi: 10.1093/g3journal/jkad256. G3 (Bethesda). 2023. PMID: 37950891 Free PMC article.
References
-
- Angers, B., K. Gharbi and A. Estoup, 2002. Evidence of gene conversion events between paralogous sequences produced by tetraploidization in Salmoninae fish. J. Mol. Evol. 54: 501–510. - PubMed
-
- Backstrom, N., H. Ceplitis, S. Berlin and H. Ellegren, 2005. Gene conversion drives the evolution of HINTW, an ampliconic gene on the female-specific avian W chromosome. Mol. Biol. Evol. 22: 1992–1999. - PubMed
-
- Brink, R., and D. Styles, 1966. A collection of pericarp factors. Maize Genet. Coop. Newsl. 40: 149–160.
-
- Carroll, S. B., J. K. Grenier and S. D. Weatherbee, 2004. From DNA to Diversity: Molecular Genetics and the Evolution of Animal Design. Blackwell Science, Malden, MA.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

