Computational expression deconvolution in a complex mammalian organ
- PMID: 16817968
- PMCID: PMC1559723
- DOI: 10.1186/1471-2105-7-328
Computational expression deconvolution in a complex mammalian organ
Abstract
Background: Microarray expression profiling has been widely used to identify differentially expressed genes in complex cellular systems. However, while such methods can be used to directly infer intracellular regulation within homogeneous cell populations, interpretation of in vivo gene expression data derived from complex organs composed of multiple cell types is more problematic. Specifically, observed changes in gene expression may be due either to changes in gene regulation within a given cell type or to changes in the relative abundance of expressing cell types. Consequently, bona fide changes in intrinsic gene regulation may be either mimicked or masked by changes in the relative proportion of different cell types. To date, few analytical approaches have addressed this problem.
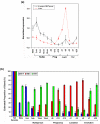
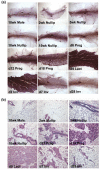
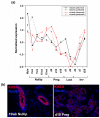
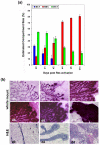
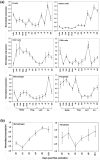
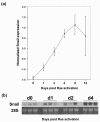
Results: We have chosen to apply a computational method for deconvoluting gene expression profiles derived from intact tissues by using reference expression data for purified populations of the constituent cell types of the mammary gland. These data were used to estimate changes in the relative proportions of different cell types during murine mammary gland development and Ras-induced mammary tumorigenesis. These computational estimates of changing compartment sizes were then used to enrich lists of differentially expressed genes for transcripts that change as a function of intrinsic intracellular regulation rather than shifts in the relative abundance of expressing cell types. Using this approach, we have demonstrated that adjusting mammary gene expression profiles for changes in three principal compartments--epithelium, white adipose tissue, and brown adipose tissue--is sufficient both to reduce false-positive changes in gene expression due solely to changes in compartment sizes and to reduce false-negative changes by unmasking genuine alterations in gene expression that were otherwise obscured by changes in compartment sizes.
Conclusion: By adjusting gene expression values for changes in the sizes of cell type-specific compartments, this computational deconvolution method has the potential to increase both the sensitivity and specificity of differential gene expression experiments performed on complex tissues. Given the necessity for understanding complex biological processes such as development and carcinogenesis within the context of intact tissues, this approach offers substantial utility and should be broadly applicable to identifying gene expression changes in tissues composed of multiple cell types.
Figures

Similar articles
-
Transcriptome analysis of mammary epithelial subpopulations identifies novel determinants of lineage commitment and cell fate.BMC Genomics. 2008 Dec 8;9:591. doi: 10.1186/1471-2164-9-591. BMC Genomics. 2008. PMID: 19063729 Free PMC article.
-
A novel doxycycline-inducible system for the transgenic analysis of mammary gland biology.FASEB J. 2002 Mar;16(3):283-92. doi: 10.1096/fj.01-0551com. FASEB J. 2002. PMID: 11874978
-
Genomic analysis of early murine mammary gland development using novel probe-level algorithms.Genome Biol. 2005;6(2):R20. doi: 10.1186/gb-2005-6-2-r20. Epub 2005 Feb 1. Genome Biol. 2005. PMID: 15693949 Free PMC article.
-
Microarray analysis of the involution switch.J Mammary Gland Biol Neoplasia. 2003 Jul;8(3):309-19. doi: 10.1023/b:jomg.0000010031.53310.92. J Mammary Gland Biol Neoplasia. 2003. PMID: 14973375 Review.
-
Computational deconvolution of transcriptomics data from mixed cell populations.Bioinformatics. 2018 Jun 1;34(11):1969-1979. doi: 10.1093/bioinformatics/bty019. Bioinformatics. 2018. PMID: 29351586 Review.
Cited by
-
MicroRNA dysregulation in the spinal cord following traumatic injury.PLoS One. 2012;7(4):e34534. doi: 10.1371/journal.pone.0034534. Epub 2012 Apr 12. PLoS One. 2012. PMID: 22511948 Free PMC article.
-
The brown adipocyte differentiation pathway in birds: an evolutionary road not taken.BMC Biol. 2008 Apr 21;6:17. doi: 10.1186/1741-7007-6-17. BMC Biol. 2008. PMID: 18426587 Free PMC article.
-
Data-driven detection of subtype-specific differentially expressed genes.Sci Rep. 2021 Jan 11;11(1):332. doi: 10.1038/s41598-020-79704-1. Sci Rep. 2021. PMID: 33432005 Free PMC article.
-
Digital sorting of complex tissues for cell type-specific gene expression profiles.BMC Bioinformatics. 2013 Mar 7;14:89. doi: 10.1186/1471-2105-14-89. BMC Bioinformatics. 2013. PMID: 23497278 Free PMC article.
-
Systematic bias in genomic classification due to contaminating non-neoplastic tissue in breast tumor samples.BMC Med Genomics. 2011 Jun 30;4:54. doi: 10.1186/1755-8794-4-54. BMC Med Genomics. 2011. PMID: 21718502 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

