The two Escherichia coli chromosome arms locate to separate cell halves
- PMID: 16818605
- PMCID: PMC1522069
- DOI: 10.1101/gad.388406
The two Escherichia coli chromosome arms locate to separate cell halves
Abstract
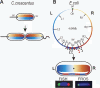
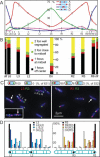
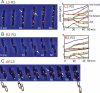
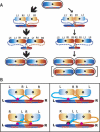
DNA replication divides the circular Escherichia coli chromosome into equal arms (replichores). Visualization of pairwise combinations of multiple genetic loci reveals that the two replichores occupy separate nucleoid halves, with the replication origin between; positions of loci on each replichore recapitulate the genetic map. Sequential replication-segregation regenerates the <left-right> structure by sequentially layering newly replicated replichore DNA to specific inner and outer edges of the developing sister nucleoids. Replication fork-dependent locus positions are imprinted, so that in most generations the <left-right> chromosome orientation in a mother cell is recreated as a <left-right-left-right> arrangement of sister chromosomes in daughter cells.
Figures
Similar articles
-
Dancing around the divisome: asymmetric chromosome segregation in Escherichia coli.Genes Dev. 2005 Oct 1;19(19):2367-77. doi: 10.1101/gad.345305. Genes Dev. 2005. PMID: 16204186 Free PMC article.
-
The Escherichia coli chromosome is organized with the left and right chromosome arms in separate cell halves.Mol Microbiol. 2006 Oct;62(2):331-8. doi: 10.1111/j.1365-2958.2006.05346.x. Mol Microbiol. 2006. PMID: 17020576
-
Replication-directed sister chromosome alignment in Escherichia coli.Mol Microbiol. 2010 Mar;75(5):1090-7. doi: 10.1111/j.1365-2958.2009.06791.x. Mol Microbiol. 2010. PMID: 20487299 Free PMC article.
-
Structural and physical aspects of bacterial chromosome segregation.J Struct Biol. 2006 Nov;156(2):273-83. doi: 10.1016/j.jsb.2006.04.013. Epub 2006 May 20. J Struct Biol. 2006. PMID: 16828313 Review.
-
The replication fork trap and termination of chromosome replication.Mol Microbiol. 2008 Dec;70(6):1323-33. doi: 10.1111/j.1365-2958.2008.06500.x. Epub 2008 Oct 17. Mol Microbiol. 2008. PMID: 19019156 Review.
Cited by
-
Insensitivity of chromosome I and the cell cycle to blockage of replication and segregation of Vibrio cholerae chromosome II.mBio. 2012 May 8;3(3):e00067-12. doi: 10.1128/mBio.00067-12. Print 2012. mBio. 2012. PMID: 22570276 Free PMC article.
-
Cell morphology and nucleoid dynamics in dividing Deinococcus radiodurans.Nat Commun. 2019 Aug 23;10(1):3815. doi: 10.1038/s41467-019-11725-5. Nat Commun. 2019. PMID: 31444361 Free PMC article.
-
How to get (a)round: mechanisms controlling growth and division of coccoid bacteria.Nat Rev Microbiol. 2013 Sep;11(9):601-14. doi: 10.1038/nrmicro3088. Nat Rev Microbiol. 2013. PMID: 23949602 Review.
-
Asymmetry of chromosome Replichores renders the DNA translocase activity of FtsK essential for cell division and cell shape maintenance in Escherichia coli.PLoS Genet. 2008 Dec;4(12):e1000288. doi: 10.1371/journal.pgen.1000288. Epub 2008 Dec 5. PLoS Genet. 2008. PMID: 19057667 Free PMC article.
-
Rules and Exceptions: The Role of Chromosomal ParB in DNA Segregation and Other Cellular Processes.Microorganisms. 2020 Jan 11;8(1):0. doi: 10.3390/microorganisms8010105. Microorganisms. 2020. PMID: 31940850 Free PMC article. Review.
References
-
- Adachi S., Kohiyama M., Onogi T., Hiraga S. Localization of replication forks in wild-type and mukB mutant cells of. Escherichia coli. Mol. Genet. Genomics. 2005;274:264–271. - PubMed
-
- Blakely G., Colloms S., May G., Burke M., Sherratt D. Escherichia coli XerC recombinase is required for chromosomal segregation at cell division. New Biol. 1991;3:789–798. - PubMed
-
- Blattner F.R., Plunkett G., III, Bloch C.A., Perna N.T., Burland V., Riley M., Collado-Vides J., Glasner J.D., Rode C.K., Mayhew G.F., et al. The complete genome sequence of Escherichia coli K-12. Science. 1997;277:1453–1474. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
