Markov propagation of allosteric effects in biomolecular systems: application to GroEL-GroES
- PMID: 16820777
- PMCID: PMC1681507
- DOI: 10.1038/msb4100075
Markov propagation of allosteric effects in biomolecular systems: application to GroEL-GroES
Abstract
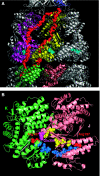
We introduce a novel approach for elucidating the potential pathways of allosteric communication in biomolecular systems. The methodology, based on Markov propagation of 'information' across the structure, permits us to partition the network of interactions into soft clusters distinguished by their coherent stochastics. Probabilistic participation of residues in these clusters defines the communication patterns inherent to the network architecture. Application to bacterial chaperonin complex GroEL-GroES, an allostery-driven structure, identifies residues engaged in intra- and inter-subunit communication, including those acting as hubs and messengers. A number of residues are distinguished by their high potentials to transmit allosteric signals, including Pro33 and Thr90 at the nucleotide-binding site and Glu461 and Arg197 mediating inter- and intra-ring communication, respectively. We propose two most likely pathways of signal transmission, between nucleotide- and GroES-binding sites across the cis and trans rings, which involve several conserved residues. A striking observation is the opposite direction of information flow within cis and trans rings, consistent with negative inter-ring cooperativity. Comparison with collective modes deduced from normal mode analysis reveals the propensity of global hinge regions to act as messengers in the transmission of allosteric signals.
Figures
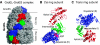





Similar articles
-
Allostery wiring diagrams in the transitions that drive the GroEL reaction cycle.J Mol Biol. 2009 Mar 27;387(2):390-406. doi: 10.1016/j.jmb.2008.12.032. Epub 2008 Dec 24. J Mol Biol. 2009. PMID: 19121324
-
Mapping pathways of allosteric communication in GroEL by analysis of correlated mutations.Proteins. 2002 Sep 1;48(4):611-7. doi: 10.1002/prot.10180. Proteins. 2002. PMID: 12211028
-
Markov methods for hierarchical coarse-graining of large protein dynamics.J Comput Biol. 2007 Jul-Aug;14(6):765-76. doi: 10.1089/cmb.2007.R015. J Comput Biol. 2007. PMID: 17691893
-
The GroEL-GroES Chaperonin Machine: A Nano-Cage for Protein Folding.Trends Biochem Sci. 2016 Jan;41(1):62-76. doi: 10.1016/j.tibs.2015.07.009. Epub 2015 Sep 25. Trends Biochem Sci. 2016. PMID: 26422689 Review.
-
GroEL/GroES: structure and function of a two-stroke folding machine.J Struct Biol. 1998 Dec 15;124(2-3):129-41. doi: 10.1006/jsbi.1998.4060. J Struct Biol. 1998. PMID: 10049801 Review.
Cited by
-
Mapping allosteric communications within individual proteins.Nat Commun. 2020 Jul 31;11(1):3862. doi: 10.1038/s41467-020-17618-2. Nat Commun. 2020. PMID: 32737291 Free PMC article.
-
Allosteric transitions in the chaperonin GroEL are captured by a dominant normal mode that is most robust to sequence variations.Biophys J. 2007 Oct 1;93(7):2289-99. doi: 10.1529/biophysj.107.105270. Epub 2007 Jun 8. Biophys J. 2007. PMID: 17557788 Free PMC article.
-
Combining structural and coevolution information to unveil allosteric sites.Chem Sci. 2023 Jun 8;14(25):7057-7067. doi: 10.1039/d2sc06272k. eCollection 2023 Jun 28. Chem Sci. 2023. PMID: 37389247 Free PMC article.
-
Directional Force Originating from ATP Hydrolysis Drives the GroEL Conformational Change.Biophys J. 2017 Apr 25;112(8):1561-1570. doi: 10.1016/j.bpj.2017.03.004. Biophys J. 2017. PMID: 28445748 Free PMC article.
-
Comparative Study of Elastic Network Model and Protein Contact Network for Protein Complexes: The Hemoglobin Case.Biomed Res Int. 2017;2017:2483264. doi: 10.1155/2017/2483264. Epub 2017 Jan 22. Biomed Res Int. 2017. PMID: 28243596 Free PMC article.
References
-
- Aharoni A, Horovitz A (1996) Inter-ring communication is disrupted in the GroEL mutant Arg13 → Gly; Ala126 → Val with known crystal structure. J Mol Biol 258: 732–735 - PubMed
-
- Bahar I, Erman B, Monnerie L (1994) Effect of molecular structures on local chain dynamics: analytical approaches and computational methods. Adv Polym Sci 116: 151–206
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

