Submicroscopic deletion in patients with Williams-Beuren syndrome influences expression levels of the nonhemizygous flanking genes
- PMID: 16826523
- PMCID: PMC1559497
- DOI: 10.1086/506371
Submicroscopic deletion in patients with Williams-Beuren syndrome influences expression levels of the nonhemizygous flanking genes
Abstract
Genomic imbalance is a common cause of phenotypic abnormalities. We measured the relative expression level of genes that map within the microdeletion that causes Williams-Beuren syndrome and within its flanking regions. We found, unexpectedly, that not only hemizygous genes but also normal-copy neighboring genes show decreased relative levels of expression. Our results suggest that not only the aneuploid genes but also the flanking genes that map several megabases away from a genomic rearrangement should be considered possible contributors to the phenotypic variation in genomic disorders.
Figures

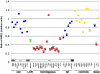

References
Web Resources
-
- Coriell Institute for Medical Research, http://www.coriell.org/index.php/content/view/31/78/ (for cell lines)
-
- Galliera Genetic Bank, http://ggb.galliera.it (for cell lines)
-
- geNorm, http://medgen.ugent.be/~jvdesomp/genorm/ (for selection of normalization genes)
-
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/ (for WBS) - PubMed
-
- UniGene, http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=unigene (for sets of transcript sequences that appear to come from the same transcription locus)
References
-
- Sebat J, Lakshmi B, Troge J, Alexander J, Young J, Lundin P, Maner S, Massa H, Walker M, Chi M, Navin N, Lucito R, Healy J, Hicks J, Ye K, Reiner A, Gilliam TC, Trask B, Patterson N, Zetterberg A, Wigler M (2004) Large-scale copy number polymorphism in the human genome. Science 305:525–52810.1126/science.1098918 - DOI - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

