Pitx2 regulates cardiac left-right asymmetry by patterning second cardiac lineage-derived myocardium
- PMID: 16836994
- PMCID: PMC5851592
- DOI: 10.1016/j.ydbio.2006.06.009
Pitx2 regulates cardiac left-right asymmetry by patterning second cardiac lineage-derived myocardium
Abstract
Current models of left-right asymmetry hold that an early asymmetric signal is generated at the node and transduced to lateral plate mesoderm in a linear signal transduction cascade through the function of the Nodal signaling molecule. The Pitx2 homeobox gene functions at the final stages of this cascade to direct asymmetric morphogenesis of selected organs including the heart. We previously showed that Pitx2 regulated an asymmetric pathway that was independent of cardiac looping suggesting a second asymmetric cardiac pathway. It has been proposed that in the cardiac outflow tract Pitx2 functions in both cardiac neural crest, as a target of canonical Wnt-signaling, and in the mesoderm-derived cardiac second lineage. We used fate mapping, conditional loss of function, and chimera analysis in mice to investigate the role of Pitx2 in outflow tract morphogenesis. Our findings reveal that Pitx2 is dispensable in the cardiac neural crest but functions in second lineage myocardium revealing that this cardiac progenitor field is patterned asymmetrically.
Figures

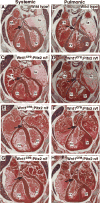


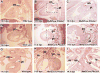


References
-
- Bajolle F, Zaffran S, Kelly RG, Hadchouel J, Bonnet D, Brown NA, Buckingham ME. Rotation of the myocardial wall of the outflow tract is implicated in the normal positioning of the great arteries. Circ Res. 2006;98:421–428. - PubMed
-
- Bostrom MP, Hutchins GM. Arrested rotation of the outflow tract may explain double-outlet right ventricle. Circulation. 1988;77:1258–1265. - PubMed
-
- Brault V, Moore R, Kutsch S, Ishibashi M, Rowitch DH, McMahon AP, Sommer L, Boussadia O, Kemler R. Inactivation of the beta-catenin gene by Wnt1-Cre-mediated deletion results in dramatic brain malformation and failure of craniofacial development. Development. 2001;128:1253–1264. - PubMed
-
- Brown NA, Anderson RH. Symmetry and laterality in the human heart: developmental implications. In: Harvey RP, Rosenthal N, editors. Heart Development. Academic Press; San Diego, CA: 1999. pp. 447–462.
-
- Brown NA, Wolpert L. The development of handedness in left/right asymmetry. Development. 1990;109:1–9. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

