Surface mutagenesis of the bovine papillomavirus E1 DNA binding domain reveals residues required for multiple functions related to DNA replication
- PMID: 16840329
- PMCID: PMC1563737
- DOI: 10.1128/JVI.00435-06
Surface mutagenesis of the bovine papillomavirus E1 DNA binding domain reveals residues required for multiple functions related to DNA replication
Abstract
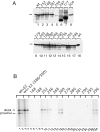
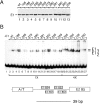
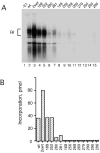
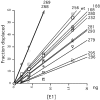
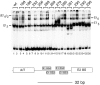
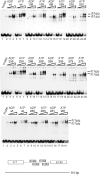
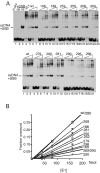
The E1 protein from papillomaviruses is a multifunctional protein with complex functions required for the initiation of viral DNA replication. We have performed a surface mutagenesis of the well-characterized E1 DNA binding domain (DBD). We demonstrate that substitutions of multiple residues on the surface of the E1 DBD are defective for DNA replication without affecting the DNA binding activity of the protein. The defects of individual substitutions include failure to form the double trimer that melts the ori and failure to form the double hexamer that unwinds the ori. These results demonstrate that the DBD plays an essential role in multiple DNA replication-related processes apart from DNA binding.
Figures

Similar articles
-
Characterization of the DNA-binding domain of the bovine papillomavirus replication initiator E1.J Virol. 1998 Apr;72(4):2567-76. doi: 10.1128/JVI.72.4.2567-2576.1998. J Virol. 1998. PMID: 9525573 Free PMC article.
-
Two patches of amino acids on the E2 DNA binding domain define the surface for interaction with E1.J Virol. 2000 Feb;74(3):1506-12. doi: 10.1128/jvi.74.3.1506-1512.2000. J Virol. 2000. PMID: 10627562 Free PMC article.
-
Binding of the E1 and E2 proteins to the origin of replication of bovine papillomavirus.J Virol. 1997 Apr;71(4):2887-96. doi: 10.1128/JVI.71.4.2887-2896.1997. J Virol. 1997. PMID: 9060646 Free PMC article.
-
Papillomavirus E1 proteins: form, function, and features.Virus Genes. 2002 Jun;24(3):275-90. doi: 10.1023/a:1015336817836. Virus Genes. 2002. PMID: 12086149 Review.
-
Papillomavirus DNA replication.Intervirology. 1994;37(3-4):150-8. doi: 10.1159/000150373. Intervirology. 1994. PMID: 7843926 Review. No abstract available.
Cited by
-
Mechanisms of Viral DNA Replication of Human Papillomavirus: E2 Protein-Dependent Recruitment of E1 DNA Helicase to the Origin of DNA Replication.Int J Mol Sci. 2025 May 2;26(9):4333. doi: 10.3390/ijms26094333. Int J Mol Sci. 2025. PMID: 40362569 Free PMC article.
-
The papillomavirus E2 proteins.Virology. 2013 Oct;445(1-2):57-79. doi: 10.1016/j.virol.2013.06.006. Epub 2013 Jul 10. Virology. 2013. PMID: 23849793 Free PMC article. Review.
-
ATP-dependent minor groove recognition of TA base pairs is required for template melting by the E1 initiator protein.J Virol. 2007 Apr;81(7):3293-302. doi: 10.1128/JVI.02432-06. Epub 2007 Jan 3. J Virol. 2007. PMID: 17202221 Free PMC article.
-
The E1 proteins.Virology. 2013 Oct;445(1-2):35-56. doi: 10.1016/j.virol.2013.07.020. Epub 2013 Sep 10. Virology. 2013. PMID: 24029589 Free PMC article. Review.
-
'Modulation of the enzymatic activities of replicative helicase (DnaB) by interaction with Hp0897: a possible mechanism for helicase loading in Helicobacter pylori'.Nucleic Acids Res. 2016 Apr 20;44(7):3288-303. doi: 10.1093/nar/gkw148. Epub 2016 Mar 21. Nucleic Acids Res. 2016. PMID: 27001508 Free PMC article.
References
-
- Bullock, P. A. 1997. The initiation of simian virus 40 DNA replication in vitro. Crit. Rev. Biochem. Mol. Biol. 32:503-568. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

