metaSHARK: a WWW platform for interactive exploration of metabolic networks
- PMID: 16845107
- PMCID: PMC1538829
- DOI: 10.1093/nar/gkl196
metaSHARK: a WWW platform for interactive exploration of metabolic networks
Abstract
The metaSHARK (metabolic search and reconstruction kit) web server offers users an intuitive, fully interactive way to explore the KEGG metabolic network via a WWW browser. Metabolic reconstruction information for specific organisms, produced by our automated SHARKhunt tool or from other programs or genome annotations, may be uploaded to the website and overlaid on the generic network. Additional data from gene expression experiments can also be incorporated, allowing the visualization of differential gene expression in the context of the predicted metabolic network. metaSHARK is available at http://bioinformatics.leeds.ac.uk/shark/.
Figures
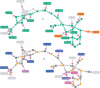
Similar articles
-
Integration of metabolic networks and gene expression in virtual reality.Bioinformatics. 2005 Sep 15;21(18):3645-50. doi: 10.1093/bioinformatics/bti581. Epub 2005 Jul 14. Bioinformatics. 2005. PMID: 16020466
-
MetaPath Online: a web server implementation of the network expansion algorithm.Nucleic Acids Res. 2007 Jul;35(Web Server issue):W613-8. doi: 10.1093/nar/gkm287. Epub 2007 May 5. Nucleic Acids Res. 2007. PMID: 17483511 Free PMC article.
-
AISMIG--an interactive server-side molecule image generator.Nucleic Acids Res. 2005 Jul 1;33(Web Server issue):W705-9. doi: 10.1093/nar/gki438. Nucleic Acids Res. 2005. PMID: 15980568 Free PMC article.
-
PathExpress: a web-based tool to identify relevant pathways in gene expression data.Nucleic Acids Res. 2007 Jul;35(Web Server issue):W176-81. doi: 10.1093/nar/gkm261. Epub 2007 Jun 22. Nucleic Acids Res. 2007. PMID: 17586825 Free PMC article.
-
Bringing Molecular Dynamics Simulation Data into View.Trends Biochem Sci. 2019 Nov;44(11):902-913. doi: 10.1016/j.tibs.2019.06.004. Epub 2019 Jul 10. Trends Biochem Sci. 2019. PMID: 31301982 Review.
Cited by
-
Model-driven analysis of experimentally determined growth phenotypes for 465 yeast gene deletion mutants under 16 different conditions.Genome Biol. 2008;9(9):R140. doi: 10.1186/gb-2008-9-9-r140. Epub 2008 Sep 22. Genome Biol. 2008. PMID: 18808699 Free PMC article.
-
A unique dual activity amino acid hydroxylase in Toxoplasma gondii.PLoS One. 2009;4(3):e4801. doi: 10.1371/journal.pone.0004801. Epub 2009 Mar 11. PLoS One. 2009. PMID: 19277211 Free PMC article.
-
A survey of visualization tools for biological network analysis.BioData Min. 2008 Nov 28;1:12. doi: 10.1186/1756-0381-1-12. BioData Min. 2008. PMID: 19040716 Free PMC article.
-
Computational meta'omics for microbial community studies.Mol Syst Biol. 2013 May 14;9:666. doi: 10.1038/msb.2013.22. Mol Syst Biol. 2013. PMID: 23670539 Free PMC article. Review.
-
Translational web robots for pathogen genome analysis.Microb Inform Exp. 2011 Oct 31;1(1):10. doi: 10.1186/2042-5783-1-10. Microb Inform Exp. 2011. PMID: 22587672 Free PMC article. No abstract available.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

