Evaluation of multiprotein immunoaffinity subtraction for plasma proteomics and candidate biomarker discovery using mass spectrometry
- PMID: 16854842
- PMCID: PMC1850944
- DOI: 10.1074/mcp.T600039-MCP200
Evaluation of multiprotein immunoaffinity subtraction for plasma proteomics and candidate biomarker discovery using mass spectrometry
Abstract
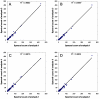
Strategies for removal of high abundance proteins have been increasingly utilized in proteomic studies of serum/plasma and other body fluids to enhance the detection of low abundance proteins and achieve broader proteome coverage; however, both the reproducibility and specificity of the high abundance protein depletion process still represent common concerns. Here we report a detailed evaluation of immunoaffinity subtraction performed applying the ProteomeLab IgY-12 system that is commonly used in human serum/plasma proteome characterization in combination with high resolution LC-MS/MS. Plasma samples were repeatedly processed using this approach, and the resulting flow-through fractions and bound fractions were individually analyzed for comparison. The removal of target proteins by the immunoaffinity subtraction system and the overall process was highly reproducible. Non-target proteins, including one spiked protein standard (rabbit glyceraldehyde-3-phosphate dehydrogenase), were also observed to bind to the column at different levels but also in a reproducible manner. The results suggest that multiprotein immunoaffinity subtraction systems can be readily integrated into quantitative strategies to enhance detection of low abundance proteins in biomarker discovery studies.
Figures
Similar articles
-
Limitation of immunoaffinity column for the removal of abundant proteins from plasma in quantitative plasma proteomics.Biomed Chromatogr. 2009 May;23(5):480-7. doi: 10.1002/bmc.1139. Biomed Chromatogr. 2009. PMID: 19039805
-
Mouse-specific tandem IgY7-SuperMix immunoaffinity separations for improved LC-MS/MS coverage of the plasma proteome.J Proteome Res. 2009 Nov;8(11):5387-95. doi: 10.1021/pr900564f. J Proteome Res. 2009. PMID: 19722698 Free PMC article.
-
Multi-component immunoaffinity subtraction chromatography: an innovative step towards a comprehensive survey of the human plasma proteome.Proteomics. 2003 Apr;3(4):422-32. doi: 10.1002/pmic.200390057. Proteomics. 2003. PMID: 12687610
-
IgY14 and SuperMix immunoaffinity separations coupled with liquid chromatography-mass spectrometry for human plasma proteomics biomarker discovery.Methods. 2012 Feb;56(2):246-53. doi: 10.1016/j.ymeth.2011.09.001. Epub 2011 Sep 10. Methods. 2012. PMID: 21925605 Free PMC article. Review.
-
Emerging Affinity-Based Proteomic Technologies for Large-Scale Plasma Profiling in Cardiovascular Disease.Circulation. 2017 Apr 25;135(17):1651-1664. doi: 10.1161/CIRCULATIONAHA.116.025446. Circulation. 2017. PMID: 28438806 Free PMC article. Review.
Cited by
-
Targeted quantification of low ng/mL level proteins in human serum without immunoaffinity depletion.J Proteome Res. 2013 Jul 5;12(7):3353-61. doi: 10.1021/pr400178v. Epub 2013 Jun 13. J Proteome Res. 2013. PMID: 23763644 Free PMC article.
-
Differential profiling of breast cancer plasma proteome by isotope-coded affinity tagging method reveals biotinidase as a breast cancer biomarker.BMC Cancer. 2010 Mar 26;10:114. doi: 10.1186/1471-2407-10-114. BMC Cancer. 2010. PMID: 20346108 Free PMC article.
-
Advances and challenges in liquid chromatography-mass spectrometry-based proteomics profiling for clinical applications.Mol Cell Proteomics. 2006 Oct;5(10):1727-44. doi: 10.1074/mcp.M600162-MCP200. Epub 2006 Aug 3. Mol Cell Proteomics. 2006. PMID: 16887931 Free PMC article. Review.
-
Emerging affinity-based techniques in proteomics.Expert Rev Proteomics. 2009 Oct;6(5):573-83. doi: 10.1586/epr.09.74. Expert Rev Proteomics. 2009. PMID: 19811078 Free PMC article. Review.
-
Proteomics studies reveal important information on small molecule therapeutics: a case study on plasma proteins.Drug Discov Today. 2008 Dec;13(23-24):1042-51. doi: 10.1016/j.drudis.2008.09.013. Epub 2008 Nov 7. Drug Discov Today. 2008. PMID: 18973825 Free PMC article. Review.
References
-
- Anderson NL, Anderson NG. The human plasma proteome: history, character, and diagnostic prospects. Mol. Cell. Proteomics. 2002;1:845–867. - PubMed
-
- Pieper R, Su Q, Gatlin CL, Huang ST, Anderson NL, Steiner S. Multi-component immunoaffinity subtraction chromatography: an innovative step towards a comprehensive survey of the human plasma proteome. Proteomics. 2003;3:422–432. - PubMed
-
- Zolotarjova N, Martosella J, Nicol G, Bailey J, Boyes BE, Barrett WC. Differences among techniques for high-abundant protein depletion. Proteomics. 2005;5:3304–3313. - PubMed
-
- Huang L, Harvie G, Feitelson JS, Gramatikoff K, Herold DA, Allen DL, Amunngama R, Hagler RA, Pisano MR, Zhang WW, Fang X. Immunoaffinity separation of plasma proteins by IgY microbeads: meeting the needs of proteomic sample preparation and analysis. Proteomics. 2005;5:3314–3328. - PubMed
-
- Chromy BA, Gonzales AD, Perkins J, Choi MW, Corzett MH, Chang BC, Corzett CH, McCutchen-Maloney SL. Proteomic analysis of human serum by two-dimensional differential gel electrophoresis after depletion of high-abundant proteins. J. Proteome Res. 2004;3:1120–1127. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials