Deciphering past human population movements in Oceania: provably optimal trees of 127 mtDNA genomes
- PMID: 16855009
- PMCID: PMC2674580
- DOI: 10.1093/molbev/msl063
Deciphering past human population movements in Oceania: provably optimal trees of 127 mtDNA genomes
Abstract
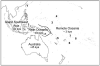
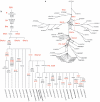
The settlement of the many island groups of Remote Oceania occurred relatively late in prehistory, beginning approximately 3,000 years ago when people sailed eastwards into the Pacific from Near Oceania, where evidence of human settlement dates from as early as 40,000 years ago. Archeological and linguistic analyses have suggested the settlers of Remote Oceania had ancestry in Taiwan, as descendants of a proposed Neolithic expansion that began approximately 5,500 years ago. Other researchers have suggested that the settlers were descendants of peoples from Island Southeast Asia or the existing inhabitants of Near Oceania alone. To explore patterns of maternal descent in Oceania, we have assembled and analyzed a data set of 137 mitochondrial DNA (mtDNA) genomes from Oceania, Australia, Island Southeast Asia, and Taiwan that includes 19 sequences generated for this project. Using the MinMax Squeeze Approach (MMS), we report the consensus network of 165 most parsimonious trees for the Oceanic data set, increasing by many orders of magnitude the numbers of trees for which a provable minimal solution has been found. The new mtDNA sequences highlight the limitations of partial sequencing for assigning sequences to haplogroups and dating recent divergence events. The provably optimal trees found for the entire mtDNA sequences using the MMS method provide a reliable and robust framework for the interpretation of evolutionary relationships and confirm that the female settlers of Remote Oceania descended from both the existing inhabitants of Near Oceania and more recent migrants into the region.
Figures

Similar articles
-
Ancient migration routes of Austronesian-speaking populations in oceanic Southeast Asia and Melanesia might mimic the spread of nasopharyngeal carcinoma.Chin J Cancer. 2011 Feb;30(2):96-105. doi: 10.5732/cjc.010.10589. Chin J Cancer. 2011. PMID: 21272441 Free PMC article. Review.
-
Phylogeny and ancient DNA of Sus provides insights into neolithic expansion in Island Southeast Asia and Oceania.Proc Natl Acad Sci U S A. 2007 Mar 20;104(12):4834-9. doi: 10.1073/pnas.0607753104. Epub 2007 Mar 14. Proc Natl Acad Sci U S A. 2007. PMID: 17360400 Free PMC article.
-
Bridging near and remote Oceania: mtDNA and NRY variation in the Solomon Islands.Mol Biol Evol. 2012 Feb;29(2):545-64. doi: 10.1093/molbev/msr186. Epub 2011 Jul 18. Mol Biol Evol. 2012. PMID: 21771715
-
Maternal history of Oceania from complete mtDNA genomes: contrasting ancient diversity with recent homogenization due to the Austronesian expansion.Am J Hum Genet. 2014 May 1;94(5):721-33. doi: 10.1016/j.ajhg.2014.03.014. Epub 2014 Apr 10. Am J Hum Genet. 2014. PMID: 24726474 Free PMC article.
-
Recent developments in the genetic history of East Asia and Oceania.Curr Opin Genet Dev. 2014 Dec;29:9-14. doi: 10.1016/j.gde.2014.06.010. Epub 2014 Aug 23. Curr Opin Genet Dev. 2014. PMID: 25170982 Review.
Cited by
-
A mitochondrial stratigraphy for island southeast Asia.Am J Hum Genet. 2007 Jan;80(1):29-43. doi: 10.1086/510412. Epub 2006 Nov 16. Am J Hum Genet. 2007. PMID: 17160892 Free PMC article.
-
Melanesian mtDNA complexity.PLoS One. 2007 Feb 28;2(2):e248. doi: 10.1371/journal.pone.0000248. PLoS One. 2007. PMID: 17327912 Free PMC article.
-
Ancient migration routes of Austronesian-speaking populations in oceanic Southeast Asia and Melanesia might mimic the spread of nasopharyngeal carcinoma.Chin J Cancer. 2011 Feb;30(2):96-105. doi: 10.5732/cjc.010.10589. Chin J Cancer. 2011. PMID: 21272441 Free PMC article. Review.
-
Mitochondrial diversity within modern human populations.Nucleic Acids Res. 2007;35(9):3039-45. doi: 10.1093/nar/gkm207. Epub 2007 Apr 16. Nucleic Acids Res. 2007. PMID: 17439969 Free PMC article.
-
Ancient voyaging and Polynesian origins.Am J Hum Genet. 2011 Feb 11;88(2):239-47. doi: 10.1016/j.ajhg.2011.01.009. Epub 2011 Feb 3. Am J Hum Genet. 2011. PMID: 21295281 Free PMC article.
References
-
- Andrews RM, Kubacka I, Chinnery PF, Lightowlers RN, Turnbull DM, Howell N. Reanalysis and revision of the Cambridge reference sequence for human mitochondrial DNA. Nat Genet. 1999;23:147. - PubMed
-
- Bellwood P. The Austronesian dispersal and the origin of languages. Sci Am. 1991;265:70–5.
-
- Bellwood P. Early agriculturalist population diasporas? Farming, languages, and genes. Annu Rev Anthropol. 2001;30:181–207.
-
- Cox MP. Indonesian mitochondrial DNA and its opposition to a Pleistocene era origin of proto-Polynesians in island southeast Asia. Hum Biol. 2005;77:179–88. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

