Diverse flavonoids stimulate NodD1 binding to nod gene promoters in Sinorhizobium meliloti
- PMID: 16855231
- PMCID: PMC1540014
- DOI: 10.1128/JB.00376-06
Diverse flavonoids stimulate NodD1 binding to nod gene promoters in Sinorhizobium meliloti
Abstract
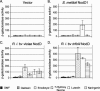
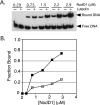
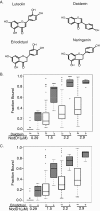
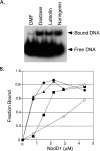
NodD1 is a member of the NodD family of LysR-type transcriptional regulators that mediates the expression of nodulation (nod) genes in the soil bacterium Sinorhizobium meliloti. Each species of rhizobia establishes a symbiosis with a limited set of leguminous plants. This host specificity results in part from a NodD-dependent upregulation of nod genes in response to a cocktail of flavonoids in the host plant's root exudates. To demonstrate that NodD is a key determinant of host specificity, we expressed nodD genes from different species of rhizobia in a strain of S. meliloti lacking endogenous NodD activity. We observed that nod gene expression was initiated in response to distinct sets of flavonoid inducers depending on the source of NodD. To better understand the effects of flavonoids on NodD, we assayed the DNA binding activity of S. meliloti NodD1 treated with the flavonoid inducer luteolin. In the presence of luteolin, NodD1 exhibited increased binding to nod gene promoters compared to binding in the absence of luteolin. Surprisingly, although they do not stimulate nod gene expression in S. meliloti, the flavonoids naringenin, eriodictyol, and daidzein also stimulated an increase in the DNA binding affinity of NodD1 to nod gene promoters. In vivo competition assays demonstrate that noninducing flavonoids act as competitive inhibitors of luteolin, suggesting that both inducing and noninducing flavonoids are able to directly bind to NodD1 and mediate conformational changes at nod gene promoters but that only luteolin is capable of promoting the downstream changes necessary for nod gene induction.
Figures


References
-
- Adhya, S., M. Gottesman, S. Garges, and A. Oppenheim. 1993. Promoter resurrection by activators—a minireview. Gene 132:1-6. - PubMed
-
- Akakura, R., and S. C. Winans. 2002. Mutations in the occQ operator that decrease OccR-induced DNA bending do not cause constitutive promoter activity. J. Biol. Chem. 277:15773-15780. - PubMed
-
- Banfalvi, Z., A. Nieuwkoop, M. Schell, L. Besl, and G. Stacey. 1988. Regulation of nod gene expression in Bradyrhizobium japonicum. Mol. Gen. Genet. 214:420-424. - PubMed
-
- Bender, G. L., M. Nayudu, K. K. Le Strange, and B. G. Rolfe. 1988. The nodD1 gene from Rhizobium strain NGR234 is a key determinant in the extension of host range to the nonlegume Parasponia. Mol. Plant-Microbe Interact. 1:259-266.
-
- Beringer, J. E. 1974. R factor transfer in Rhizobium leguminosarum. J. Gen. Microbiol. 84:188-198. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

