Mutational analysis of the promoter recognized by Chlamydia and Escherichia coli sigma(28) RNA polymerase
- PMID: 16855242
- PMCID: PMC1540034
- DOI: 10.1128/JB.00480-06
Mutational analysis of the promoter recognized by Chlamydia and Escherichia coli sigma(28) RNA polymerase
Abstract
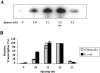
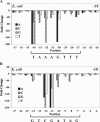
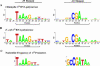
sigma(28) RNA polymerase is an alternative RNA polymerase that has been postulated to have a role in developmental gene regulation in Chlamydia. Although a consensus bacterial sigma(28) promoter sequence has been proposed, it is based on a relatively small number of defined promoters, and the promoter structure has not been systematically analyzed. To evaluate the sequence of the sigma(28)-dependent promoter, we performed a comprehensive mutational analysis of the Chlamydia trachomatis hctB promoter, testing the effect of point substitutions on promoter activity. We defined a -35 element recognized by chlamydial sigma(28) RNA polymerase that resembles the consensus -35 sequence. Within the -10 element, however, chlamydial sigma(28) RNA polymerase showed a striking preference for a CGA sequence at positions -12 to -10 rather than the longer consensus -10 sequence. We also observed a strong preference for this CGA sequence by Escherichia coli sigma(28) RNA polymerase, suggesting that this previously unrecognized motif is the critical component of the -10 promoter element recognized by sigma(28) RNA polymerase. Although the consensus spacer length is 11 nucleotides (nt), we found that sigma(28) RNA polymerase from both Chlamydia and E. coli transcribed a promoter with either an 11- or 12-nt spacer equally well. Altogether, we found very similar results for sigma(28) RNA polymerase from C. trachomatis and E. coli, suggesting that promoter recognition by this alternative RNA polymerase is well conserved among bacteria. The preferred sigma(28) promoter that we defined in the context of the hctB promoter is TAAAGwwy-n(11/12)-ryCGAwrn, where w is A or T, r is a purine, y is a pyrimidine, n is any nucleotide, and n(11/12) is a spacer of 11 or 12 nt.
Figures
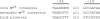

Similar articles
-
Mutational analysis of the Chlamydia trachomatis dnaK promoter defines the optimal -35 promoter element.Nucleic Acids Res. 2003 Jan 15;31(2):551-5. doi: 10.1093/nar/gkg150. Nucleic Acids Res. 2003. PMID: 12527761 Free PMC article.
-
Mutagenesis of region 4 of sigma 28 from Chlamydia trachomatis defines determinants for protein-protein and protein-DNA interactions.J Bacteriol. 2009 Jan;191(2):651-60. doi: 10.1128/JB.01083-08. Epub 2008 Oct 31. J Bacteriol. 2009. PMID: 18978051 Free PMC article.
-
Chlamydia trachomatis RNA polymerase major sigma subunit. Sequence and structural comparison of conserved and unique regions with Escherichia coli sigma 70 and Bacillus subtilis sigma 43.J Biol Chem. 1990 Aug 5;265(22):13206-14. J Biol Chem. 1990. PMID: 2142944
-
Mutational analysis of structure-function relationship of RNA polymerase in Escherichia coli.Methods Enzymol. 1996;273:300-19. doi: 10.1016/s0076-6879(96)73027-6. Methods Enzymol. 1996. PMID: 8791620 Review. No abstract available.
-
[The molecular anatomy of RNA polymerase].Seikagaku. 1995 Feb;67(2):123-30. Seikagaku. 1995. PMID: 7759911 Review. Japanese. No abstract available.
Cited by
-
The growing repertoire of genetic tools for dissecting chlamydial pathogenesis.Pathog Dis. 2021 May 11;79(5):ftab025. doi: 10.1093/femspd/ftab025. Pathog Dis. 2021. PMID: 33930127 Free PMC article.
-
Promoter analysis of macrophage- and tick cell-specific differentially expressed Ehrlichia chaffeensis p28-Omp genes.BMC Microbiol. 2009 May 19;9:99. doi: 10.1186/1471-2180-9-99. BMC Microbiol. 2009. PMID: 19454021 Free PMC article.
-
A novel short L-arginine responsive protein-coding gene (laoB) antiparallel overlapping to a CadC-like transcriptional regulator in Escherichia coli O157:H7 Sakai originated by overprinting.BMC Evol Biol. 2018 Feb 12;18(1):21. doi: 10.1186/s12862-018-1134-0. BMC Evol Biol. 2018. PMID: 29433444 Free PMC article.
-
The transcriptional repressor EUO regulates both subsets of Chlamydia late genes.Mol Microbiol. 2014 Nov;94(4):888-97. doi: 10.1111/mmi.12804. Epub 2014 Oct 16. Mol Microbiol. 2014. PMID: 25250726 Free PMC article.
-
The latent cis-regulatory potential of mobile DNA in Escherichia coli.Nat Commun. 2025 May 21;16(1):4740. doi: 10.1038/s41467-025-60023-w. Nat Commun. 2025. PMID: 40399339 Free PMC article.
References
-
- Chesnokova, O., J. B. Coutinho, I. H. Khan, M. S. Mikhail, and C. I. Kado. 1997. Characterization of flagella genes of Agrobacterium tumefaciens, and the effect of a bald strain on virulence. Mol. Microbiol. 23:579-590. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

