Reevaluation of Drosophila melanogaster's neuronal circadian pacemakers reveals new neuronal classes
- PMID: 16856134
- PMCID: PMC2596765
- DOI: 10.1002/cne.21021
Reevaluation of Drosophila melanogaster's neuronal circadian pacemakers reveals new neuronal classes
Abstract
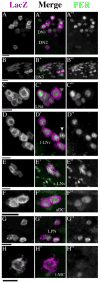
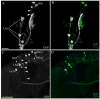
In the brain of the fly Drosophila melanogaster, approximately 150 clock-neurons are organized to synchronize and maintain behavioral rhythms, but the physiological and neurochemical bases of their interactions are largely unknown. Here we reevaluate the cellular properties of these pacemakers by application of a novel genetic reporter and several phenotypic markers. First, we describe an enhancer trap marker called R32 that specifically reveals several previously undescribed aspects of the fly's central neuronal pacemakers. We find evidence for a previously unappreciated class of neuronal pacemakers, the lateral posterior neurons (LPNs), and establish anatomical, molecular, and developmental criteria to establish a subclass within the dorsal neuron 1 (DN1) group of pacemakers. Furthermore, we show that the neuropeptide IPNamide is specifically expressed by this DN1 subclass. These observations implicate IPNamide as a second candidate circadian transmitter in the Drosophila brain. Finally, we present molecular and anatomical evidence for unrecognized phenotypic diversity within each of four established classes of clock neurons.
2006 Wiley-Liss, Inc.
Figures








Similar articles
-
Electrical silencing of PDF neurons advances the phase of non-PDF clock neurons in Drosophila.J Biol Rhythms. 2008 Apr;23(2):117-28. doi: 10.1177/0748730407312984. J Biol Rhythms. 2008. PMID: 18375861
-
The Drosophila circadian clock is a variably coupled network of multiple peptidergic units.Science. 2014 Mar 28;343(6178):1516-20. doi: 10.1126/science.1251285. Science. 2014. PMID: 24675961 Free PMC article.
-
A Screening of UNF Targets Identifies Rnb, a Novel Regulator of Drosophila Circadian Rhythms.J Neurosci. 2017 Jul 12;37(28):6673-6685. doi: 10.1523/JNEUROSCI.3286-16.2017. Epub 2017 Jun 7. J Neurosci. 2017. PMID: 28592698 Free PMC article.
-
Neurobiology of the fruit fly's circadian clock.Genes Brain Behav. 2005 Mar;4(2):65-76. doi: 10.1111/j.1601-183X.2004.00092.x. Genes Brain Behav. 2005. PMID: 15720403 Review.
-
Organization of the Drosophila circadian control circuit.Curr Biol. 2008 Jan 22;18(2):R84-93. doi: 10.1016/j.cub.2007.11.061. Curr Biol. 2008. PMID: 18211849 Review.
Cited by
-
Moonlight shifts the endogenous clock of Drosophila melanogaster.Proc Natl Acad Sci U S A. 2007 Feb 27;104(9):3538-43. doi: 10.1073/pnas.0606870104. Epub 2007 Feb 16. Proc Natl Acad Sci U S A. 2007. PMID: 17307880 Free PMC article.
-
The neuropeptide PDF acts directly on evening pacemaker neurons to regulate multiple features of circadian behavior.PLoS Biol. 2009 Jul;7(7):e1000154. doi: 10.1371/journal.pbio.1000154. Epub 2009 Jul 21. PLoS Biol. 2009. PMID: 19621061 Free PMC article.
-
Circadian pacemaker neurons change synaptic contacts across the day.Curr Biol. 2014 Sep 22;24(18):2161-2167. doi: 10.1016/j.cub.2014.07.063. Epub 2014 Aug 21. Curr Biol. 2014. PMID: 25155512 Free PMC article.
-
Circadian control of membrane excitability in Drosophila melanogaster lateral ventral clock neurons.J Neurosci. 2008 Jun 18;28(25):6493-501. doi: 10.1523/JNEUROSCI.1503-08.2008. J Neurosci. 2008. PMID: 18562620 Free PMC article.
-
Role of Tau Protein in Remodeling of Circadian Neuronal Circuits and Sleep.Front Aging Neurosci. 2019 Nov 21;11:320. doi: 10.3389/fnagi.2019.00320. eCollection 2019. Front Aging Neurosci. 2019. PMID: 31824299 Free PMC article.
References
-
- Baggerman G, Cerstiaens A, De Loof A, Schoofs L. Peptidomics of the larval Drosophila melanogaster central nervous system. J Biol Chem. 2002;277:40368–40374. - PubMed
-
- Cyran SA, Buchsbaum AM, Reddy KL, Lin MC, Glossop MR, Hardin PE, Young MW, Storti RV, Blau J. vrille, Pdp1, and dClock form a second feedback loop in the Drosophila circadian clock. Cell. 2003;112:329–341. - PubMed
-
- Dunlap JC. Molecular bases for circadian clocks. Cell. 1999;96:271–790. - PubMed
-
- Frisch B, Hardin PE, Hamblen-Coyle MJ, Rosbash M, Hall JC. A promoterless period gene mediates behavioral rhythmicity and cyclical per expression in a restricted subset of the Drosophila nervous system. Neuron. 1994;12:555–570. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

