The complexity of the mammalian transcriptome
- PMID: 16857706
- PMCID: PMC1819450
- DOI: 10.1113/jphysiol.2006.115568
The complexity of the mammalian transcriptome
Abstract
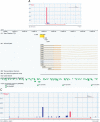
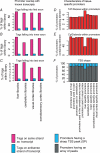
A comprehensive understanding of protein and regulatory networks is strictly dependent on the complete description of the transcriptome of cells. After the determination of the genome sequence of several mammalian species, gene identification is based on in silico predictions followed by evidence of transcription. Conservative estimates suggest that there are about 20,000 protein-encoding genes in the mammalian genome. In the last few years the combination of full-length cDNA cloning, cap-analysis gene expression (CAGE) tag sequencing and tiling arrays experiments have unveiled unexpected additional complexities in the transcriptome. Here we describe the current view of the mammalian transcriptome focusing on transcripts diversity, the growing non-coding RNA world, the organization of transcriptional units in the genome and promoter structures. In-depth analysis of the brain transcriptome has been challenging due to the cellular complexity of this organ. Here we present a computational analysis of CAGE data from different regions of the central nervous system, suggesting distinctive mechanisms of brain-specific transcription.
Figures



References
-
- Audic S, Claverie JM. The significance of digital gene expression profiles. Genome Res. 1997;7:986–995. - PubMed
-
- Bertone P, Stolc V, Royce TE, Rozowsky JS, Urban AE, Zhu X, et al. Global identification of human transcribed sequences with genome tiling arrays. Science. 2004;306:2242–2246. - PubMed
-
- Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC, Maeda N, et al. FANTOM Consortium; RIKEN Genome Exploration Research Group and Genome Science Group (Genome Network Project Core Group) The transcriptional landscape of the mammalian genome. Science. 2005;309:1559–1563. - PubMed
-
- Carninci P, Sandelin A, Lenhard B, Katayama S, Shimokawa K, Ponjavic J, et al. Genome-wide analysis of mammalian promoter architecture and evolution. Nat Genet. 2006;38:626–635. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

