The prokaryotic antecedents of the ubiquitin-signaling system and the early evolution of ubiquitin-like beta-grasp domains
- PMID: 16859499
- PMCID: PMC1779556
- DOI: 10.1186/gb-2006-7-7-r60
The prokaryotic antecedents of the ubiquitin-signaling system and the early evolution of ubiquitin-like beta-grasp domains
Abstract
Background: Ubiquitin (Ub)-mediated signaling is one of the hallmarks of all eukaryotes. Prokaryotic homologs of Ub (ThiS and MoaD) and E1 ligases have been studied in relation to sulfur incorporation reactions in thiamine and molybdenum/tungsten cofactor biosynthesis. However, there is no evidence for entire protein modification systems with Ub-like proteins and deconjugation by deubiquitinating enzymes in prokaryotes. Hence, the evolutionary assembly of the eukaryotic Ub-signaling apparatus remains unclear.
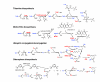
Results: We systematically analyzed prokaryotic Ub-related beta-grasp fold proteins using sensitive sequence profile searches and structural analysis. Consequently, we identified novel Ub-related proteins beyond the characterized ThiS, MoaD, TGS, and YukD domains. To understand their functional associations, we sought and recovered several conserved gene neighborhoods and domain architectures. These included novel associations involving diverse sulfur metabolism proteins, siderophore biosynthesis and the gene encoding the transfer mRNA binding protein SmpB, as well as domain fusions between Ub-like domains and PIN-domain related RNAses. Most strikingly, we found conserved gene neighborhoods in phylogenetically diverse bacteria combining genes for JAB domains (the primary de-ubiquitinating isopeptidases of the proteasomal complex), along with E1-like adenylating enzymes and different Ub-related proteins. Further sequence analysis of other conserved genes in these neighborhoods revealed several Ub-conjugating enzyme/E2-ligase related proteins. Genes for an Ub-like protein and a JAB domain peptidase were also found in the tail assembly gene cluster of certain caudate bacteriophages.
Conclusion: These observations imply that members of the Ub family had already formed strong functional associations with E1-like proteins, UBC/E2-related proteins, and JAB peptidases in the bacteria. Several of these Ub-like proteins and the associated protein families are likely to function together in signaling systems just as in eukaryotes.
Figures





Similar articles
-
Structure and evolution of ubiquitin and ubiquitin-related domains.Methods Mol Biol. 2012;832:15-63. doi: 10.1007/978-1-61779-474-2_2. Methods Mol Biol. 2012. PMID: 22350875
-
Functional diversification of the RING finger and other binuclear treble clef domains in prokaryotes and the early evolution of the ubiquitin system.Mol Biosyst. 2011 Jul;7(7):2261-77. doi: 10.1039/c1mb05061c. Epub 2011 May 6. Mol Biosyst. 2011. PMID: 21547297 Free PMC article.
-
Novel predicted peptidases with a potential role in the ubiquitin signaling pathway.Cell Cycle. 2004 Nov;3(11):1440-50. doi: 10.4161/cc.3.11.1206. Epub 2004 Nov 6. Cell Cycle. 2004. PMID: 15483401
-
The natural history of ubiquitin and ubiquitin-related domains.Front Biosci (Landmark Ed). 2012 Jan 1;17(4):1433-60. doi: 10.2741/3996. Front Biosci (Landmark Ed). 2012. PMID: 22201813 Free PMC article. Review.
-
Prokaryotic ubiquitin-like protein modification.Annu Rev Microbiol. 2014;68:155-75. doi: 10.1146/annurev-micro-091313-103447. Epub 2014 May 29. Annu Rev Microbiol. 2014. PMID: 24995873 Free PMC article. Review.
Cited by
-
A conserved signaling pathway activates bacterial CBASS immune signaling in response to DNA damage.EMBO J. 2022 Nov 17;41(22):e111540. doi: 10.15252/embj.2022111540. Epub 2022 Sep 26. EMBO J. 2022. PMID: 36156805 Free PMC article.
-
The Anti-sigma Factor RsiV Is a Bacterial Receptor for Lysozyme: Co-crystal Structure Determination and Demonstration That Binding of Lysozyme to RsiV Is Required for σV Activation.PLoS Genet. 2016 Sep 7;12(9):e1006287. doi: 10.1371/journal.pgen.1006287. eCollection 2016 Sep. PLoS Genet. 2016. PMID: 27602573 Free PMC article.
-
Impact of whole-genome and tandem duplications in the expansion and functional diversification of the F-box family in legumes (Fabaceae).PLoS One. 2013;8(2):e55127. doi: 10.1371/journal.pone.0055127. Epub 2013 Feb 4. PLoS One. 2013. PMID: 23390519 Free PMC article.
-
The Ubiquitin Moiety of Ubi1 Is Required for Productive Expression of Ribosomal Protein eL40 in Saccharomyces cerevisiae.Cells. 2019 Aug 7;8(8):850. doi: 10.3390/cells8080850. Cells. 2019. PMID: 31394841 Free PMC article.
-
Role of the ubiquitin-like protein Urm1 as a noncanonical lysine-directed protein modifier.Proc Natl Acad Sci U S A. 2011 Feb 1;108(5):1763-70. doi: 10.1073/pnas.1014402108. Epub 2011 Jan 5. Proc Natl Acad Sci U S A. 2011. PMID: 21209336 Free PMC article.
References
-
- Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P. Molecular Biology of the Cell, (book and CD-ROM) 4. New York, NY: Garland Science Publishing; 2002.
-
- Ardley HC, Robinson PA. E3 ubiquitin ligases. Essays Biochem. 2005;41:15–30. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

