Signal transduction, gene transcription, and cytokine production triggered in macrophages by exposure to trypanosome DNA
- PMID: 16861639
- PMCID: PMC1539588
- DOI: 10.1128/IAI.01938-05
Signal transduction, gene transcription, and cytokine production triggered in macrophages by exposure to trypanosome DNA
Abstract
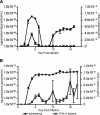
Activation of a type I cytokine response is important for early resistance to infection with Trypanosoma brucei rhodesiense, the extracellular protozoan parasite that causes African sleeping sickness. The work presented here demonstrates that trypanosome DNA activates macrophages to produce factors that may contribute to this response. Initial results demonstrated that T. brucei rhodesiense DNA was present in the plasma of C57BL/6 and C57BL/6-scid mice following infection. Subsequently, the effect of trypanosome DNA on macrophages was investigated; parasite DNA was found to be less stimulatory than Escherichia coli DNA but more stimulatory than murine DNA, as predicted by the CG dinucleotide content. Trypanosome DNA stimulated the induction of a signal transduction cascade associated with Toll-like receptor signaling in RAW 264.7 macrophage cells. The signaling cascade led to expression of mRNAs, including interleukin-12 (IL-12) p40, IL-6, IL-10, cyclooxygenase-2, and beta interferon. The treatment of RAW 264.7 cells and bone marrow-derived macrophages with trypanosome DNA induced the production of NO, prostaglandin E2, and the cytokines IL-6, IL-10, IL-12, and tumor necrosis factor alpha. In all cases, DNase I treatment of T. brucei rhodesisense DNA abolished the activation. These results suggest that T. brucei rhodesiense DNA serves as a ligand for innate immune cells and may play an important contributory role in early stimulation of the host immune response during trypanosomiasis.
Figures

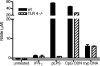



Similar articles
-
Innate resistance to experimental African trypanosomiasis: differences in cytokine (TNF-alpha, IL-6, IL-10 and IL-12) production by bone marrow-derived macrophages from resistant and susceptible mice.Cytokine. 2000 Jul;12(7):1024-34. doi: 10.1006/cyto.2000.0685. Cytokine. 2000. PMID: 10880248
-
Analysis of macrophage activation in African trypanosomiasis.J Leukoc Biol. 2001 May;69(5):685-90. J Leukoc Biol. 2001. PMID: 11358974 Review.
-
[Human African trypanosomiasis].C R Seances Soc Biol Fil. 1996;190(4):395-408. C R Seances Soc Biol Fil. 1996. PMID: 8952890 Review. French.
-
Alternative versus classical macrophage activation during experimental African trypanosomosis.Int J Parasitol. 2001 May 1;31(5-6):575-87. doi: 10.1016/s0020-7519(01)00170-9. Int J Parasitol. 2001. PMID: 11334945 Review.
-
Glycosylinositolphosphate soluble variant surface glycoprotein inhibits IFN-gamma-induced nitric oxide production via reduction in STAT1 phosphorylation in African trypanosomiasis.J Immunol. 2003 Aug 1;171(3):1466-72. doi: 10.4049/jimmunol.171.3.1466. J Immunol. 2003. PMID: 12874239
Cited by
-
Host cell signalling and leishmania mechanisms of evasion.J Trop Med. 2012;2012:819512. doi: 10.1155/2012/819512. Epub 2011 Nov 3. J Trop Med. 2012. PMID: 22131998 Free PMC article.
-
Urinary tract infections caused by staphylococcus aureus DNA in comparison to the candida albicans DNA.N Am J Med Sci. 2011 Dec;3(12):565-9. doi: 10.4297/najms.2011.3562. N Am J Med Sci. 2011. PMID: 22363080 Free PMC article.
-
Salivarian Trypanosomosis: A Review of Parasites Involved, Their Global Distribution and Their Interaction With the Innate and Adaptive Mammalian Host Immune System.Front Immunol. 2018 Oct 2;9:2253. doi: 10.3389/fimmu.2018.02253. eCollection 2018. Front Immunol. 2018. PMID: 30333827 Free PMC article. Review.
-
Type I IFNs play a role in early resistance, but subsequent susceptibility, to the African trypanosomes.J Immunol. 2008 Oct 1;181(7):4908-17. doi: 10.4049/jimmunol.181.7.4908. J Immunol. 2008. PMID: 18802094 Free PMC article.
-
The biocompatibility of sulfobetaine engineered polymethylmethacrylate by surface entrapment technique.J Mater Sci Mater Med. 2010 Feb;21(2):635-46. doi: 10.1007/s10856-009-3886-y. Epub 2009 Oct 11. J Mater Sci Mater Med. 2010. PMID: 19821070
References
-
- Akira, S., and K. Takeda. 2004. Toll-like receptor signalling. Nat. Rev. Immunol. 4:499-511. - PubMed
-
- Baetselier, P. D., B. Namangala, W. Noel, L. Brys, E. Pays, and A. Beschin. 2001. Alternative versus classical macrophage activation during experimental African trypanosomosis. Int. J. Parasitol. 31:575-587. - PubMed
-
- Coller, S. P., J. M. Mansfield, and D. M. Paulnock. 2003. Glycosylinositolphosphate soluble variant surface glycoprotein inhibits IFN-gamma-induced nitric oxide production via reduction in STAT1 phosphorylation in African trypanosomiasis. J. Immunol. 171:1466-1472. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

