Improved in situ hybridization efficiency with locked-nucleic-acid-incorporated DNA probes
- PMID: 16885281
- PMCID: PMC1538721
- DOI: 10.1128/AEM.03039-05
Improved in situ hybridization efficiency with locked-nucleic-acid-incorporated DNA probes
Abstract
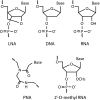
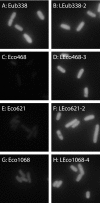
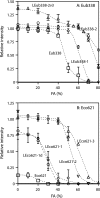
Low signal intensity due to poor probe hybridization efficiency is one of the major drawbacks of rRNA-targeted in situ hybridization. There are two major factors affecting the hybridization efficiency: probe accessibility and affinity to the targeted rRNA molecules. In this study, we demonstrate remarkable improvement in in situ hybridization efficiency by applying locked-nucleic-acid (LNA)-incorporated oligodeoxynucleotide probes (LNA/DNA probes) without compromising specificity. Fluorescently labeled LNA/DNA probes with two to four LNA substitutions exhibited strong fluorescence intensities equal to or greater than that of probe Eub338, although these probes did not show bright signals when they were synthesized as DNA probes; for example, the fluorescence intensity of probe Eco468 increased by 22-fold after three LNA bases were substituted for DNA bases. Dissociation profiles of the probes revealed that the dissociation temperature was directly related to the number of LNA substitutions and the fluorescence intensity. These results suggest that the introduction of LNA residues in DNA probes will be a useful approach for effectively enhancing probe hybridization efficiency.
Figures

Similar articles
-
Dramatically improved RNA in situ hybridization signals using LNA-modified probes.RNA. 2005 Nov;11(11):1745-8. doi: 10.1261/rna.2139705. Epub 2005 Sep 21. RNA. 2005. PMID: 16177135 Free PMC article.
-
LNA-modified oligonucleotides are highly efficient as FISH probes.Cytogenet Genome Res. 2004;107(1-2):32-7. doi: 10.1159/000079569. Cytogenet Genome Res. 2004. PMID: 15305054
-
Synthesis and investigation of deoxyribonucleic acid/locked nucleic acid chimeric molecular beacons.Nucleic Acids Res. 2007;35(12):4030-41. doi: 10.1093/nar/gkm358. Epub 2007 Jun 8. Nucleic Acids Res. 2007. PMID: 17557813 Free PMC article.
-
LNA (locked nucleic acid): high-affinity targeting of complementary RNA and DNA.Biochemistry. 2004 Oct 26;43(42):13233-41. doi: 10.1021/bi0485732. Biochemistry. 2004. PMID: 15491130 Review.
-
LNA: a versatile tool for therapeutics and genomics.Trends Biotechnol. 2003 Feb;21(2):74-81. doi: 10.1016/S0167-7799(02)00038-0. Trends Biotechnol. 2003. PMID: 12573856 Review.
Cited by
-
Demonstration of hepatitis C virus RNA with in situ hybridization employing a locked nucleic Acid probe in humanized liver of infected chimeric mice and in needle-biopsied human liver.Int J Hepatol. 2013;2013:249535. doi: 10.1155/2013/249535. Epub 2013 Jun 18. Int J Hepatol. 2013. PMID: 23853723 Free PMC article.
-
LNA probes substantially improve the detection of bacterial endosymbionts in whole mount of insects by fluorescent in-situ hybridization.BMC Microbiol. 2012 May 24;12:81. doi: 10.1186/1471-2180-12-81. BMC Microbiol. 2012. PMID: 22624773 Free PMC article.
-
Heterogeneity of miR-10b expression in circulating tumor cells.Sci Rep. 2015 Nov 2;5:15980. doi: 10.1038/srep15980. Sci Rep. 2015. PMID: 26522916 Free PMC article.
-
Evaluation of fluorescence in situ hybridization techniques to study long non-coding RNA expression in cultured cells.Nucleic Acids Res. 2018 Jan 9;46(1):e4. doi: 10.1093/nar/gkx946. Nucleic Acids Res. 2018. PMID: 29059327 Free PMC article.
-
Unlocked nucleic acid modified primer-based enzymatic polymerization assay: towards allele-specific genotype detection of human platelet antigens.RSC Adv. 2018 Sep 21;8(57):32770-32774. doi: 10.1039/c8ra06050a. eCollection 2018 Sep 18. RSC Adv. 2018. PMID: 35547719 Free PMC article.
References
-
- Amann, R. I., B. M. Fuchs, and S. Behrens. 2001. The identification of microorganisms by fluorescence in situ hybridisation. Curr. Opin. Biotechnol. 12:231-236. - PubMed
-
- Behrens, S., C. Rühland, J. Inácio, H. Huber, Á. Fonseca, I. Spencer-Martins, B. M. Fuchs, and R. Amann. 2003. In situ accessibility of small-subunit rRNA of members of the domains Bacteria, Archaea, and Eucarya to Cy3-labeled oligonucleotide probe. Appl. Environ. Microbiol. 69:1748-1758. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

