The in vivo pattern of AID targeting to immunoglobulin switch regions deduced from mutation spectra in msh2-/- ung-/- mice
- PMID: 16894013
- PMCID: PMC2118391
- DOI: 10.1084/jem.20061067
The in vivo pattern of AID targeting to immunoglobulin switch regions deduced from mutation spectra in msh2-/- ung-/- mice
Abstract
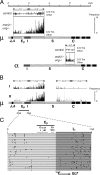
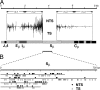
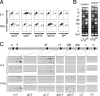
Immunoglobulin (Ig) class switching is initiated by deamination of C-->U within the immunoglobulin heavy chain locus, catalyzed by activation-induced deaminase (AID). In the absence of uracil-DNA glycosylase (UNG) and the homologue of bacterial MutS (MSH)-2 mismatch recognition protein, the resultant U:G lesions are not processed into switching events but are fixed by replication allowing sites of AID-catalyzed deamination to be identified by the resulting C-->T mutations. We find that AID targets cytosines in both donor and acceptor switch regions (S regions) with the deamination domains initiating approximately 150 nucleotides 3' of the I exon start sites and extending over several kilobases (the IgH intronic enhancer is spared). Culturing B cells with interleukin 4 or interferon gamma specifically enhanced deamination around Sgamma1 and Sgamma2a, respectively. Mutation spectra suggest that, in the absence of UNG and MSH2, AID may occasionally act at the mu switch region in an apparently processive manner, but there is no marked preference for targeting of the transcribed versus nontranscribed strand (even in areas capable of R loop formation). The data are consistent with switch recombination being triggered by transcription-associated, strand-symmetric AID-mediated deamination at both donor and acceptor S regions with cytokines directing isotype specificity by potentiating AID recruitment to the relevant acceptor S region.
Figures


References
-
- Chaudhuri, J., and F.W. Alt. 2004. Class-switch recombination: interplay of transcription, DNA deamination and DNA repair. Nat. Rev. Immunol. 4:541–552. - PubMed
-
- Kenter, A.L. 2005. Class switch recombination: an emerging mechanism. Curr. Top. Microbiol. Immunol. 290:171–199. - PubMed
-
- Longerich, S., U. Basu, F.W. Alt, and U. Storb. 2006. AID in somatic hypermutation and class switch recombination. Curr. Opin. Immunol. 18:164–174. - PubMed
-
- Stavnezer, J. 1996. Immunoglobulin class switching. Curr. Opin. Immunol. 8:199–205. - PubMed
-
- Muramatsu, M., K. Kinoshita, S. Fagarasan, S. Yamada, Y. Shinkai, and T. Honjo. 2000. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell. 102:553–563. - PubMed
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

