Consensus by democracy. Using meta-analyses of microarray and genomic data to model the cold acclimation signaling pathway in Arabidopsis
- PMID: 16896234
- PMCID: PMC1533918
- DOI: 10.1104/pp.106.083527
Consensus by democracy. Using meta-analyses of microarray and genomic data to model the cold acclimation signaling pathway in Arabidopsis
Abstract
The whole-genome response of Arabidopsis (Arabidopsis thaliana) exposed to different types and durations of abiotic stress has now been described by a wealth of publicly available microarray data. When combined with studies of how gene expression is affected in mutant and transgenic Arabidopsis with altered ability to transduce the low temperature signal, these data can be used to test the interactions between various low temperature-associated transcription factors and their regulons. We quantized a collection of Affymetrix microarray data so that each gene in a particular regulon could vote on whether a cis-element found in its promoter conferred induction (+1), repression (-1), or no transcriptional change (0) during cold stress. By statistically comparing these election results with the voting behavior of all genes on the same gene chip, we verified the bioactivity of novel cis-elements and defined whether they were inductive or repressive. Using in silico mutagenesis we identified functional binding consensus variants for the transcription factors studied. Our results suggest that the previously identified ICEr1 (induction of CBF expression region 1) consensus does not correlate with cold gene induction, while the ICEr3/ICEr4 consensuses identified using our algorithms are present in regulons of genes that were induced coordinate with observed ICE1 transcript accumulation and temporally preceding genes containing the dehydration response element. Statistical analysis of overlap and cis-element enrichment in the ICE1, CBF2, ZAT12, HOS9, and PHYA regulons enabled us to construct a regulatory network supported by multiple lines of evidence that can be used for future hypothesis testing.
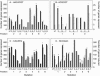
Figures




Similar articles
-
Roles of the CBF2 and ZAT12 transcription factors in configuring the low temperature transcriptome of Arabidopsis.Plant J. 2005 Jan;41(2):195-211. doi: 10.1111/j.1365-313X.2004.02288.x. Plant J. 2005. PMID: 15634197
-
An Arabidopsis homeodomain transcription factor gene, HOS9, mediates cold tolerance through a CBF-independent pathway.Proc Natl Acad Sci U S A. 2004 Jun 29;101(26):9873-8. doi: 10.1073/pnas.0403166101. Epub 2004 Jun 17. Proc Natl Acad Sci U S A. 2004. PMID: 15205481 Free PMC article.
-
The zinc-finger protein Zat12 plays a central role in reactive oxygen and abiotic stress signaling in Arabidopsis.Plant Physiol. 2005 Oct;139(2):847-56. doi: 10.1104/pp.105.068254. Epub 2005 Sep 23. Plant Physiol. 2005. PMID: 16183833 Free PMC article.
-
Gene Regulatory Networks Mediating Cold Acclimation: The CBF Pathway.Adv Exp Med Biol. 2018;1081:3-22. doi: 10.1007/978-981-13-1244-1_1. Adv Exp Med Biol. 2018. PMID: 30288701 Review.
-
CBF-dependent signaling pathway: a key responder to low temperature stress in plants.Crit Rev Biotechnol. 2011 Jun;31(2):186-92. doi: 10.3109/07388551.2010.505910. Epub 2010 Oct 4. Crit Rev Biotechnol. 2011. PMID: 20919819 Review.
Cited by
-
Global expression profiling of low temperature induced genes in the chilling tolerant japonica rice Jumli Marshi.PLoS One. 2013 Dec 12;8(12):e81729. doi: 10.1371/journal.pone.0081729. eCollection 2013. PLoS One. 2013. PMID: 24349120 Free PMC article.
-
Development of a model system to identify differences in spring and winter oat.PLoS One. 2012;7(1):e29792. doi: 10.1371/journal.pone.0029792. Epub 2012 Jan 9. PLoS One. 2012. PMID: 22253782 Free PMC article.
-
Transcriptome analysis identifies novel responses and potential regulatory genes involved in seasonal dormancy transitions of leafy spurge (Euphorbia esula L.).BMC Genomics. 2008 Nov 12;9:536. doi: 10.1186/1471-2164-9-536. BMC Genomics. 2008. PMID: 19014493 Free PMC article.
-
CsICE1 and CsCBF1: two transcription factors involved in cold responses in Camellia sinensis.Plant Cell Rep. 2012 Jan;31(1):27-34. doi: 10.1007/s00299-011-1136-5. Epub 2011 Aug 18. Plant Cell Rep. 2012. PMID: 21850593
-
Male gametophyte-specific WRKY34 transcription factor mediates cold sensitivity of mature pollen in Arabidopsis.J Exp Bot. 2010 Sep;61(14):3901-14. doi: 10.1093/jxb/erq204. Epub 2010 Jul 19. J Exp Bot. 2010. PMID: 20643804 Free PMC article.
References
-
- Boyce JM, Knight H, Deyholos M, Openshaw MR, Galbraith DW, Warren G, Knight MR (2003) The sfr6 mutant of Arabidopsis is defective in transcriptional activation via CBF/DREB1 and DREB2 and shows sensitivity to osmotic stress. Plant J 34: 395–406 - PubMed
-
- Chinnusamy V, Zhu J, Zhu J-K (2006) Gene regulation during cold acclimation in plants. Physiol Plant 126: 52–61
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

