An hpr1 point mutation that impairs transcription and mRNP biogenesis without increasing recombination
- PMID: 16908536
- PMCID: PMC1636866
- DOI: 10.1128/MCB.00684-06
An hpr1 point mutation that impairs transcription and mRNP biogenesis without increasing recombination
Abstract
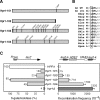
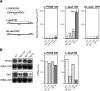
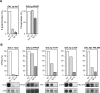
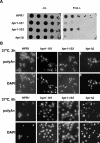
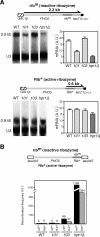
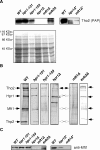
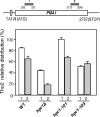
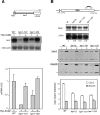
THO/TREX, a conserved eukaryotic protein complex, is a key player at the interface between transcription and mRNP metabolism. The lack of a functional THO complex impairs transcription, leads to transcription-dependent hyperrecombination, causes mRNA export defects and fast mRNA decay, and retards replication fork progression in a transcription-dependent manner. To get more insight into the interconnection between mRNP biogenesis and genomic instability, we searched for HPR1 mutations that differentially affect gene expression and recombination. We isolated mutants that were barely affected in gene expression but exhibited a hyperrecombination phenotype. In addition, we isolated a mutant, hpr1-101, with a strong defect in transcription, as observed for lacZ, and a general defect in mRNA export that did not display a relevant hyperrecombination phenotype. In THO single-null mutants, but not in the hpr1 point mutants studied, THO and its subunits were unstable. Interestingly, in contrast to hyperrecombinant null mutants, hpr1-101 did not cause retardation of replication fork progression. Transcription and mRNP biogenesis can therefore be impaired by THO/TREX dysfunction without increasing recombination, suggesting that it is possible to separate the mechanism(s) responsible for mRNA biogenesis defects from the further step of triggering transcription-dependent recombination.
Figures

References
-
- Aguilera, A. 2005. Cotranscriptional mRNP assembly: from the DNA to the nuclear pore. Curr. Opin. Cell Biol. 17:242-250. - PubMed
-
- Amberg, D. C., A. L. Goldstein, and C. N. Cole. 1992. Isolation and characterization of RAT1: an essential gene of Saccharomyces cerevisiae required for the efficient nucleocytoplasmic trafficking of mRNA. Genes Dev. 6:1173-1189. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
