Endoplasmic reticulum stress and neurodegeneration in rats neonatally infected with borna disease virus
- PMID: 16912310
- PMCID: PMC1563873
- DOI: 10.1128/JVI.00836-06
Endoplasmic reticulum stress and neurodegeneration in rats neonatally infected with borna disease virus
Abstract
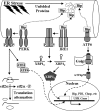
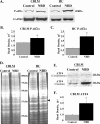
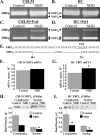
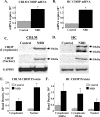
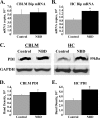
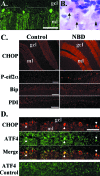
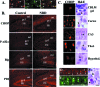
Borna disease virus infection of neonatal rats results in a characteristic behavioral syndrome and apoptosis of subsets of neurons in the hippocampus and cerebellum (neonatal Borna disease [NBD]). The cellular mechanisms leading to neurodevelopmental damage in NBD have not been fully elucidated. Insights into this model may have general implications for understanding the pathogenesis of virus-associated neurodevelopmental damage. Here we report the presence of endoplasmic reticulum (ER) stress markers and activation of the unfolded protein response in the NBD hippocampus and cerebellum. Specific findings included enhanced PERK-mediated phosphorylation of eif2alpha and concomitant regulation of ATF4 translation; IRE1-mediated splicing of XBP1 mRNA; and cleavage of the ATF6 protein in NBD rat brains. We found evidence for regional and cell type-specific divergence in the expression of ER stress-induced proapoptotic and quality control signals. Our results demonstrate that ER stress induction in death-susceptible Purkinje neurons in NBD is associated with the expression of the proapoptotic molecule CHOP in the absence of compensatory expression of the ER quality control molecules Bip and protein disulfide isomerase. In contrast, ER stress in death-resistant astrocytes is associated with complementary expression of CHOP and ER quality control signals. These results implicate an imbalance between ER stress-mediated apoptosis and survival signaling as a critical determinant of neural cell fate in NBD.
Figures


Similar articles
-
Spatiotemporal analysis of purkinje cell degeneration relative to parasagittal expression domains in a model of neonatal viral infection.J Virol. 2007 Mar;81(6):2675-87. doi: 10.1128/JVI.02245-06. Epub 2006 Dec 20. J Virol. 2007. PMID: 17182680 Free PMC article.
-
Hippocampal poly(ADP-Ribose) polymerase 1 and caspase 3 activation in neonatal bornavirus infection.J Virol. 2008 Feb;82(4):1748-58. doi: 10.1128/JVI.02014-07. Epub 2007 Dec 5. J Virol. 2008. PMID: 18057239 Free PMC article.
-
Borna disease virus replication in organotypic hippocampal slice cultures from rats results in selective damage of dentate granule cells.J Virol. 2005 Sep;79(18):11716-23. doi: 10.1128/JVI.79.18.11716-11723.2005. J Virol. 2005. PMID: 16140749 Free PMC article.
-
Borna disease virus.Rev Med Virol. 2001 Jan-Feb;11(1):37-57. doi: 10.1002/rmv.300. Rev Med Virol. 2001. PMID: 11241801 Free PMC article. Review.
-
Natural and experimental Borna disease virus infections--neuropathology and pathogenetic considerations.APMIS Suppl. 2008;(124):53-7. doi: 10.1111/j.1600-0463.2008.000m8.x. APMIS Suppl. 2008. PMID: 18771099 Review. No abstract available.
Cited by
-
ER stress in Alzheimer's disease: a novel neuronal trigger for inflammation and Alzheimer's pathology.J Neuroinflammation. 2009 Dec 26;6:41. doi: 10.1186/1742-2094-6-41. J Neuroinflammation. 2009. PMID: 20035627 Free PMC article. Review.
-
Association of Organochlorine Pesticides With Genetic Markers of Endoplasmic Reticulum Stress in Type 2 Diabetes Mellitus: A Case-Control Study Among the North-Indian Population.Front Endocrinol (Lausanne). 2022 Mar 16;13:841463. doi: 10.3389/fendo.2022.841463. eCollection 2022. Front Endocrinol (Lausanne). 2022. PMID: 35370992 Free PMC article.
-
Silencing GADD153/CHOP gene expression protects against Alzheimer's disease-like pathology induced by 27-hydroxycholesterol in rabbit hippocampus.PLoS One. 2011;6(10):e26420. doi: 10.1371/journal.pone.0026420. Epub 2011 Oct 14. PLoS One. 2011. Retraction in: PLoS One. 2024 Jul 19;19(7):e0307751. doi: 10.1371/journal.pone.0307751. PMID: 22046282 Free PMC article. Retracted.
-
Propofol induces endoplasmic reticulum (ER) stress and apoptosis in lung cancer cell H460.Tumour Biol. 2014 Jun;35(6):5213-7. doi: 10.1007/s13277-014-1677-7. Epub 2014 Feb 9. Tumour Biol. 2014. PMID: 24510348
-
HIV-1 Tat activates neuronal ryanodine receptors with rapid induction of the unfolded protein response and mitochondrial hyperpolarization.PLoS One. 2008;3(11):e3731. doi: 10.1371/journal.pone.0003731. Epub 2008 Nov 14. PLoS One. 2008. PMID: 19009018 Free PMC article.
References
-
- Barone, M. V., A. Crozat, A. Tabaee, L. Philipson, and D. Ron. 1994. CHOP (GADD153) and its oncogenic variant, TLS-CHOP, have opposing effects on the induction of G1/S arrest. Genes Dev. 8:453-464. - PubMed
-
- Bautista, J. R., S. A. Rubin, T. H. Moran, G. J. Schwartz, and K. M. Carbone. 1995. Developmental injury to the cerebellum following perinatal Borna disease virus infection. Brain Res. Dev. Brain Res. 90:45-53. - PubMed
-
- Bertolotti, A., Y. Zhang, L. M. Hendershot, H. P. Harding, and D. Ron. 2000. Dynamic interaction of BiP and ER stress transducers in the unfolded-protein response. Nat. Cell Biol. 2:326-332. - PubMed
-
- Bitko, V., and S. Barik. 2001. An endoplasmic reticulum-specific stress-activated caspase (caspase-12) is implicated in the apoptosis of A549 epithelial cells by respiratory syncytial virus. J. Cell. Biochem. 80:441-454. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

