GENCODE: producing a reference annotation for ENCODE
- PMID: 16925838
- PMCID: PMC1810553
- DOI: 10.1186/gb-2006-7-s1-s4
GENCODE: producing a reference annotation for ENCODE
Abstract
Background: The GENCODE consortium was formed to identify and map all protein-coding genes within the ENCODE regions. This was achieved by a combination of initial manual annotation by the HAVANA team, experimental validation by the GENCODE consortium and a refinement of the annotation based on these experimental results.
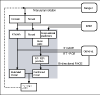
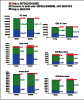
Results: The GENCODE gene features are divided into eight different categories of which only the first two (known and novel coding sequence) are confidently predicted to be protein-coding genes. 5' rapid amplification of cDNA ends (RACE) and RT-PCR were used to experimentally verify the initial annotation. Of the 420 coding loci tested, 229 RACE products have been sequenced. They supported 5' extensions of 30 loci and new splice variants in 50 loci. In addition, 46 loci without evidence for a coding sequence were validated, consisting of 31 novel and 15 putative transcripts. We assessed the comprehensiveness of the GENCODE annotation by attempting to validate all the predicted exon boundaries outside the GENCODE annotation. Out of 1,215 tested in a subset of the ENCODE regions, 14 novel exon pairs were validated, only two of them in intergenic regions.
Conclusion: In total, 487 loci, of which 434 are coding, have been annotated as part of the GENCODE reference set available from the UCSC browser. Comparison of GENCODE annotation with RefSeq and ENSEMBL show only 40% of GENCODE exons are contained within the two sets, which is a reflection of the high number of alternative splice forms with unique exons annotated. Over 50% of coding loci have been experimentally verified by 5' RACE for EGASP and the GENCODE collaboration is continuing to refine its annotation of 1% human genome with the aid of experimental validation.
Figures

References
-
- International Human Genome Sequencing Consortium Finishing the euchromatic sequence of the human genome. Nature. 2004;431:931–945. - PubMed
-
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297. - PubMed
-
- ENCODE project consortium The ENCODE (ENCyclopedia Of DNA Elements) Project. Science. 2004;306:636–640. - PubMed
-
- GENCODE Consortium http://genome.imim.es/gencode
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

