CD8 T cells control cytomegalovirus latency by epitope-specific sensing of transcriptional reactivation
- PMID: 16928768
- PMCID: PMC1641801
- DOI: 10.1128/JVI.01248-06
CD8 T cells control cytomegalovirus latency by epitope-specific sensing of transcriptional reactivation
Abstract
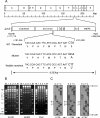
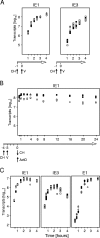
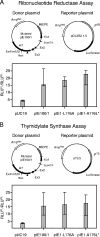
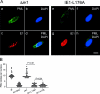
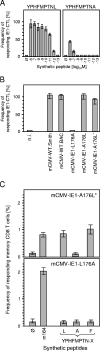
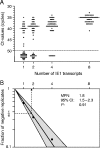
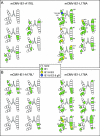
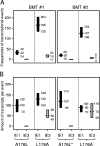
During murine cytomegalovirus (mCMV) latency in the lungs, most of the viral genomes are transcriptionally silent at the major immediate-early locus, but rare and stochastic episodes of desilencing lead to the expression of IE1 transcripts. This low-frequency but perpetual expression is accompanied by an activation of lung-resident effector-memory CD8 T cells specific for the antigenic peptide 168-YPHFMPTNL-176, which is derived from the IE1 protein. These molecular and immunological findings were combined in the "silencing/desilencing and immune sensing hypothesis" of cytomegalovirus latency and reactivation. This hypothesis proposes that IE1 gene expression proceeds to cell surface presentation of the IE1 peptide by the major histocompatibility complex (MHC) class I molecule L(d) and that its recognition by CD8 T cells terminates virus reactivation. Here we provide experimental evidence in support of this hypothesis. We generated mutant virus mCMV-IE1-L176A, in which the antigenic IE1 peptide is functionally deleted by a point mutation of the C-terminal MHC class I anchor residue Leu into Ala. Two revertant viruses, mCMV-IE1-A176L and the wobble nucleotide-marked mCMV-IE1-A176L*, in which Leu is restored by back-mutation of Ala codon GCA into Leu codons CTA and CTT, respectively, were constructed. Pulmonary latency of the mutant virus was found to be associated with an increased prevalence of IE1 transcription and with events of IE3 transactivator splicing. In conclusion, IE1-specific CD8 T cells recognize and terminate virus reactivation in vivo at the first opportunity in the reactivated gene expression program. The perpetual gene expression and antigen presentation might represent the driving molecular force in CMV-associated immunosenescence.
Figures



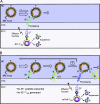
References
-
- Bain, M., M. Reeves, and J. Sinclair. 2006. Regulation of human cytomegalovirus gene expression by chromatin remodeling, p. 167-183. In M. J. Reddehase (ed.), Cytomegaloviruses: molecular biology and immunology. Caister Academic Press, Wymondham, Norfolk, United Kingdom.
-
- Bolovan-Fritts, C. A., E. S. Mocarski, and J. A. Wiedeman. 1999. Peripheral blood CD14+ cells from healthy subjects carry a circular conformation of latent cytomegalovirus genome. Blood 93:394-398. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

