Rapid and accurate pyrosequencing of angiosperm plastid genomes
- PMID: 16934154
- PMCID: PMC1564139
- DOI: 10.1186/1471-2229-6-17
Rapid and accurate pyrosequencing of angiosperm plastid genomes
Abstract
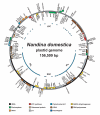
Background: Plastid genome sequence information is vital to several disciplines in plant biology, including phylogenetics and molecular biology. The past five years have witnessed a dramatic increase in the number of completely sequenced plastid genomes, fuelled largely by advances in conventional Sanger sequencing technology. Here we report a further significant reduction in time and cost for plastid genome sequencing through the successful use of a newly available pyrosequencing platform, the Genome Sequencer 20 (GS 20) System (454 Life Sciences Corporation), to rapidly and accurately sequence the whole plastid genomes of the basal eudicot angiosperms Nandina domestica (Berberidaceae) and Platanus occidentalis (Platanaceae).
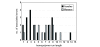
Results: More than 99.75% of each plastid genome was simultaneously obtained during two GS 20 sequence runs, to an average depth of coverage of 24.6x in Nandina and 17.3x in Platanus. The Nandina and Platanus plastid genomes shared essentially identical gene complements and possessed the typical angiosperm plastid structure and gene arrangement. To assess the accuracy of the GS 20 sequence, over 45 kilobases of sequence were generated for each genome using conventional sequencing. Overall error rates of 0.043% and 0.031% were observed in GS 20 sequence for Nandina and Platanus, respectively. More than 97% of all observed errors were associated with homopolymer runs, with approximately 60% of all errors associated with homopolymer runs of 5 or more nucleotides and approximately 50% of all errors associated with regions of extensive homopolymer runs. No substitution errors were present in either genome. Error rates were generally higher in the single-copy and noncoding regions of both plastid genomes relative to the inverted repeat and coding regions.
Conclusion: Highly accurate and essentially complete sequence information was obtained for the Nandina and Platanus plastid genomes using the GS 20 System. More importantly, the high accuracy observed in the GS 20 plastid genome sequence was generated for a significant reduction in time and cost over traditional shotgun-based genome sequencing techniques, although with approximately half the coverage of previously reported GS 20 de novo genome sequence. The GS 20 should be broadly applicable to angiosperm plastid genome sequencing, and therefore promises to expand the scale of plant genetic and phylogenetic research dramatically.
Figures


References
-
- Olmstead RG, Palmer JD. Chloroplast DNA systematics: A review of methods and data analysis. Am J Bot. 1994;81:1205–1224. doi: 10.2307/2445483. - DOI
-
- Grevich JJ, Daniell H. Chloroplast genetic engineering: Recent advances and future perspectives. Crit Rev Plant Sci. 2005;24:83–107. doi: 10.1080/07352680590935387. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

