A peroxisomal glutathione transferase of Saccharomyces cerevisiae is functionally related to sulfur amino acid metabolism
- PMID: 16936141
- PMCID: PMC1595348
- DOI: 10.1128/EC.00216-06
A peroxisomal glutathione transferase of Saccharomyces cerevisiae is functionally related to sulfur amino acid metabolism
Abstract
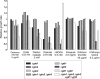
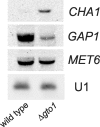
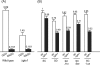
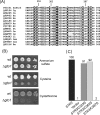
Saccharomyces cerevisiae cells contain three omega-class glutathione transferases with glutaredoxin activity (Gto1, Gto2, and Gto3), in addition to two glutathione transferases (Gtt1 and Gtt2) not classifiable into standard classes. Gto1 is located at the peroxisomes, where it is targeted through a PTS1-type sequence, whereas Gto2 and Gto3 are in the cytosol. Among the GTO genes, GTO2 shows the strongest induction of expression by agents such as diamide, 1-chloro-2,4-dinitrobenzene, tert-butyl hydroperoxide or cadmium, in a manner that is dependent on transcriptional factors Yap1 and/or Msn2/4. Diamide and 1-chloro-2,4-dinitrobenzene (causing depletion of reduced glutathione) also induce expression of GTO1 over basal levels. Phenotypic analyses with single and multiple mutants in the S. cerevisiae glutathione transferase genes show that, in the absence of Gto1 and the two Gtt proteins, cells display increased sensitivity to cadmium. A gto1-null mutant also shows growth defects on oleic acid-based medium, which is indicative of abnormal peroxisomal functions, and altered expression of genes related to sulfur amino acid metabolism. As a consequence, growth of the gto1 mutant is delayed in growth medium without lysine, serine, or threonine, and the mutant cells have low levels of reduced glutathione. The role of Gto1 at the S. cerevisiae peroxisomes could be related to the redox regulation of the Str3 cystathionine beta-lyase protein. This protein is also located at the peroxisomes in S. cerevisiae, where it is involved in transulfuration of cysteine into homocysteine, and requires a conserved cysteine residue for its biological activity.
Figures




References
-
- Adamis, P. D. B., D. S. Gomes, M. L. C. C. Pinto, A. D. Panek, and E. C. A. Eleuterio. 2004. The role of glutathione transferases in cadmium stress. Toxicol. Lett. 154:81-88. - PubMed
-
- Armstrong, R. N. 1997. Structure, catalytic mechanism, and evolution of the glutathione transferases. Chem. Res. Toxicol. 10:2-18. - PubMed
-
- Avery, S. V. 2001. Metal toxicity in yeasts and the role of oxidative stress. Adv. Appl. Microbiol. 49:111-142. - PubMed
-
- Awasthi, Y. C., G. A. S. Ansari, and S. Awasthi. 2005. Regulation of 4-hydroxynonenal mediated signaling by glutathione S-transferases. Methods Enzymol. 401:379-407. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

