Negative and positive regulation of gene expression by mouse histone deacetylase 1
- PMID: 16940178
- PMCID: PMC1636735
- DOI: 10.1128/MCB.01220-06
Negative and positive regulation of gene expression by mouse histone deacetylase 1
Abstract
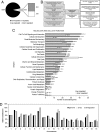
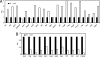
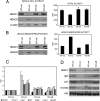
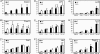
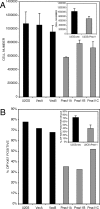
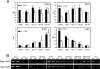
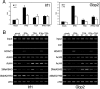
Histone deacetylases (HDACs) catalyze the removal of acetyl groups from core histones. Because of their capacity to induce local condensation of chromatin, HDACs are generally considered repressors of transcription. In this report, we analyzed the role of the class I histone deacetylase HDAC1 as a transcriptional regulator by comparing the expression profiles of wild-type and HDAC1-deficient embryonic stem cells. A specific subset of mouse genes (7%) was deregulated in the absence of HDAC1. We identified several putative tumor suppressors (JunB, Prss11, and Plagl1) and imprinted genes (Igf2, H19, and p57) as novel HDAC1 targets. The majority of HDAC1 target genes showed reduced expression accompanied by recruitment of HDAC1 and local reduction in histone acetylation at regulatory regions. At some target genes, the related deacetylase HDAC2 partially masks the loss of HDAC1. A second group of genes was found to be downregulated in HDAC1-deficient cells, predominantly by additional recruitment of HDAC2 in the absence of HDAC1. Finally, a small set of genes (Gja1, Irf1, and Gbp2) was found to require HDAC activity and recruitment of HDAC1 for their transcriptional activation. Our study reveals a regulatory cross talk between HDAC1 and HDAC2 and a novel function for HDAC1 as a transcriptional coactivator.
Figures


References
-
- Abdollahi, A., D. Pisarcik, D. Roberts, J. Weinstein, P. Cairns, and T. C. Hamilton. 2003. LOT1 (PLAGL1/ZAC1), the candidate tumor suppressor gene at chromosome 6q24-25, is epigenetically regulated in cancer. J. Biol. Chem. 278:6041-6049. - PubMed
-
- Ahringer, J. 2000. NuRD and SIN3 histone deacetylase complexes in development. Trends Genet. 16:351-356. - PubMed
-
- Baldi, A., A. De Luca, M. Morini, T. Battista, A. Felsani, F. Baldi, C. Catricala, A. Amantea, D. M. Noonan, A. Albini, P. G. Natali, D. Lombardi, and M. G. Paggi. 2002. The HtrA1 serine protease is down-regulated during human melanoma progression and represses growth of metastatic melanoma cells. Oncogene 21:6684-6688. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
