Coatomer, the coat protein of COPI transport vesicles, discriminates endoplasmic reticulum residents from p24 proteins
- PMID: 16940185
- PMCID: PMC1636745
- DOI: 10.1128/MCB.01055-06
Coatomer, the coat protein of COPI transport vesicles, discriminates endoplasmic reticulum residents from p24 proteins
Abstract
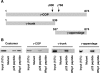
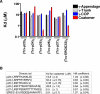
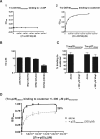
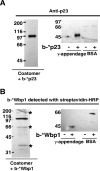
In the formation of COPI vesicles, interactions take place between the coat protein coatomer and membrane proteins: either cargo proteins for retrieval to the endoplasmic reticulum (ER) or proteins that cycle between the ER and the Golgi. While the binding sites on coatomer for ER residents have been characterized, how cycling proteins bind to the COPI coat is still not clear. In order to understand at a molecular level the mechanism of uptake of such proteins, we have investigated the binding to coatomer of p24 proteins as examples of cycling proteins as well as that of ER-resident cargos. The p24 proteins required dimerization to interact with coatomer at two independent binding sites in gamma-COP. In contrast, ER-resident cargos bind to coatomer as monomers and to sites other than gamma-COP. The COPI coat therefore discriminates between p24 proteins and ER-resident proteins by differential binding involving distinct subunits.
Figures





References
-
- Bornstein, P., and G. Balian. 1977. Cleavage at Asn-Gly bonds with hydroxylamine. Methods Enzymol. 47:132-145. - PubMed
-
- Bremser, M., W. Nickel, M. Schweikert, M. Ravazzola, M. Amherdt, C. A. Hughes, T. H. Sollner, J. E. Rothman, and F. T. Wieland. 1999. Coupling of coat assembly and vesicle budding to packaging of putative cargo receptors. Cell 96:495-506. - PubMed
-
- Contreras, I., E. Ortiz-Zapater, and F. Aniento. 2004. Sorting signals in the cytosolic tail of membrane proteins involved in the interaction with plant ARF1 and coatomer. Plant J. 38:685-698. - PubMed
-
- Cosson, P., and F. Letourneur. 1994. Coatomer interaction with di-lysine endoplasmic reticulum retention motifs. Science 263:1629-1631. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
