Molecular basis of clonal expansion of hematopoiesis in 2 patients with paroxysmal nocturnal hemoglobinuria (PNH)
- PMID: 16940417
- PMCID: PMC1895453
- DOI: 10.1182/blood-2006-05-025148
Molecular basis of clonal expansion of hematopoiesis in 2 patients with paroxysmal nocturnal hemoglobinuria (PNH)
Abstract
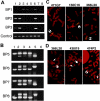
Somatic mutation of PIGA in hematopoietic stem cells causes deficiency of glycosyl phosphatidylinositol-anchored proteins in paroxysmal nocturnal hemoglobinuria (PNH) that underlies the intravascular hemolysis but does not account for expansion of the PNH clone. Immune mechanisms may mediate clonal selection but appear insufficient to account for the clonal dominance necessary for PNH to become clinically apparent. Herein, we report 2 patients with PNH whose PIGA-mutant cells had a concurrent, acquired rearrangement of chromosome 12. In both cases, der(12) had a break within the 3' untranslated region of HMGA2, the architectural transcription factor gene deregulated in many benign mesenchymal tumors, that caused ectopic expression of HMGA2 in the bone marrow. These observations suggest that aberrant HMGA2 expression, in concert with mutant PIGA, accounts for clonal hematopoiesis in these 2 patients and suggest the concept of PNH as a benign tumor of the bone marrow.
Figures
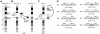

References
-
- Takeda J, Miyata T, Kawagoe K, et al. Deficiency of the GPI anchor caused by a somatic mutation of the PIG-A gene in paroxysmal nocturnal hemoglobinuria. Cell. 1993;73: 703-711. - PubMed
-
- Miyata T, Takeda J, Iida Y, et al. The cloning of PIG-A, a component in the early step of GPI-anchor biosynthesis. Science. 1993;259: 1318-1320. - PubMed
-
- Kawagoe K, Kitamura D, Okabe M, et al. Glycosylphosphatidylinositol-anchor-deficient mice: implications for clonal dominance of mutant cells in paroxysmal nocturnal hemoglobinuria. Blood. 1996;87: 3600-3606. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

