Inactivation of Rv2525c, a substrate of the twin arginine translocation (Tat) system of Mycobacterium tuberculosis, increases beta-lactam susceptibility and virulence
- PMID: 16952959
- PMCID: PMC1595485
- DOI: 10.1128/JB.00631-06
Inactivation of Rv2525c, a substrate of the twin arginine translocation (Tat) system of Mycobacterium tuberculosis, increases beta-lactam susceptibility and virulence
Abstract
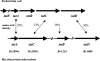
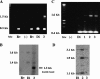
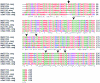
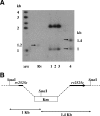
The twin arginine translocation (Tat) system is used by many bacteria to export fully folded proteins containing cofactors. Here, we show genetically that this system is essential for Mycobacterium tuberculosis, as the tatAC operon and tatB genes could be inactivated only in partially diploid strains. Using comparative genomics, the rv2525c gene of M. tuberculosis was identified as encoding a histidine-rich protein, with a twin arginine signal peptide, and orthologous genes were shown to be present in several but not all actinobacterial species. Conservation of this gene by Mycobacterium leprae, which has undergone reductive evolution, suggested an important role for rv2525c. An rv2525c knockout mutant was constructed, and biochemical analysis indicated that the mature Rv2525c protein is secreted. Upon exposure to antituberculous drugs, rv2525c expression is significantly up-regulated together with those of other genes involved in cell wall biogenesis. Phenotypic comparison of the mutant with the parental strain revealed an increase in susceptibility to some beta-lactam antibiotics and, despite slower growth in vitro, enhanced virulence in both cellular and murine models of tuberculosis. The Tat system thus contributes in multiple ways to survival of the tubercle bacillus.
Figures


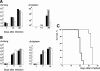
References
-
- Bange, W. R., F. M. Collins, and W. R. Jacobs, Jr. 1999. Survival of mice infected with Mycobacterium smegmatis containing large DNA fragments from Mycobacterium tuberculosis. Tuber. Lung Dis. 79:171-180. - PubMed
-
- Berks, B. C., F. Sargent, and T. Palmer. 2000. The Tat protein export pathway. Mol. Microbiol. 35:260-274. - PubMed
-
- Bogsch, E. G., F. Sargent, N. R. Stanley, B. C. Berks, C. Robinson, and T. Palmer. 1998. An essential component of a novel bacterial protein export system with homologues in plastids and mitochondria. J. Biol. Chem. 273:18003-18006. - PubMed
-
- Boshoff, H. I., T. G. Myers, B. R. Copp, M. R. McNeil, M. A. Wilson, and C. E. Barry III. 2004. The transcriptional responses of Mycobacterium tuberculosis to inhibitors of metabolism: novel insights into drug mechanisms of action. J. Biol. Chem. 279:40174-40184. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

