Real-time PCR assay for detection and quantification of hepatitis B virus genotypes A to G
- PMID: 16954268
- PMCID: PMC1594718
- DOI: 10.1128/JCM.00024-06
Real-time PCR assay for detection and quantification of hepatitis B virus genotypes A to G
Abstract
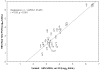
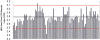
The detection and quantification of hepatitis B virus (HBV) DNA play an important role in diagnosing and monitoring HBV infection as well as assessing therapeutic response. The great variability among HBV genotypes and the enormous range of clinical HBV DNA levels present challenges for PCR-based amplification techniques. In this study, we describe the development, evaluation, and validation of a novel real-time PCR assay designed to provide accurate quantification of DNA from all eight HBV genotypes in patient plasma specimens. A computer algorithm was used to design degenerate real-time PCR primers and probes based upon a large number (n = 340) of full-length genomic sequences including HBV genotypes A to H from Europe, Africa, Asia, and North and South America. Genotype performance was tested and confirmed using 59 genotype A to G specimens from two commercially available worldwide genotype panels. This assay has a dynamic range of at least 8 log(10) without the need for specimen dilution, good clinical intra- and interassay precision, and excellent correlation with the Bayer Diagnostics VERSANT HBV DNA 3.0 (branched DNA) assay (r = 0.93). Probit analysis determined the 95% detection level was 56 IU/ml, corresponding to 11 copies per PCR well. The high sensitivity, wide linear range, good reproducibility, and genotype inclusivity, combined with a small sample volume requirement and low cost, make this novel quantitative HBV real-time PCR assay particularly well suited for application to large clinical and epidemiological studies.
Figures



Similar articles
-
Real time TaqMan PCR detection and quantitation of HBV genotypes A-G with the use of an internal quantitation standard.J Clin Virol. 2004 May;30(1):86-93. doi: 10.1016/j.jcv.2003.08.015. J Clin Virol. 2004. PMID: 15072760
-
Evaluation of the COBAS AmpliPrep-total nucleic acid isolation-COBAS TaqMan hepatitis B virus (HBV) quantitative test and comparison to the VERSANT HBV DNA 3.0 assay.J Clin Microbiol. 2006 Apr;44(4):1390-9. doi: 10.1128/JCM.44.4.1390-1399.2006. J Clin Microbiol. 2006. PMID: 16597867 Free PMC article.
-
A genotype-independent real-time PCR assay for quantification of hepatitis B virus DNA.J Clin Microbiol. 2007 Feb;45(2):553-8. doi: 10.1128/JCM.00709-06. Epub 2006 Dec 20. J Clin Microbiol. 2007. PMID: 17182753 Free PMC article.
-
Fully automated quantitation of hepatitis B virus (HBV) DNA in human plasma by the COBAS AmpliPrep/COBAS TaqMan system.J Clin Virol. 2006 Apr;35(4):373-80. doi: 10.1016/j.jcv.2006.01.003. Epub 2006 Feb 7. J Clin Virol. 2006. PMID: 16461000
-
A real-time quantitative assay for hepatitis B DNA virus (HBV) developed to detect all HBV genotypes.Rev Inst Med Trop Sao Paulo. 2010 May-Jun;52(3):119-24. doi: 10.1590/s0036-46652010000300001. Rev Inst Med Trop Sao Paulo. 2010. PMID: 20602019
Cited by
-
Molecular Detection of Epstein-Barr Virus, Human Herpes Virus 6, Cytomegalovirus, and Hepatitis B Virus in Patients with Multiple Sclerosis.Middle East J Dig Dis. 2020 Jul;12(3):171-177. doi: 10.34172/mejdd.2020.179. Middle East J Dig Dis. 2020. PMID: 33062222 Free PMC article.
-
Simple Low-Cost Production of DNA MS2 Virus-Like Particles As Molecular Diagnostic Controls.GEN Biotechnol. 2022 Dec 1;1(6):496-503. doi: 10.1089/genbio.2022.0033. Epub 2022 Dec 21. GEN Biotechnol. 2022. PMID: 36644571 Free PMC article.
-
More DNA and RNA of HBV SP1 splice variants are detected in genotypes B and C at low viral replication.Sci Rep. 2021 Dec 13;11(1):23838. doi: 10.1038/s41598-021-03304-w. Sci Rep. 2021. PMID: 34903774 Free PMC article.
-
In Vitro Lactic Acid Bacteria Anti-Hepatitis B Virus (HBV) Effect and Modulation of the Intestinal Microbiota in Fecal Cultures from HBV-Associated Hepatocellular Carcinoma Patients.Nutrients. 2024 Feb 22;16(5):600. doi: 10.3390/nu16050600. Nutrients. 2024. PMID: 38474727 Free PMC article.
-
Hepatitis B virus DNA splicing in Lebanese blood donors and genotype A to E strains: implications for hepatitis B virus DNA quantification and infectivity.J Clin Microbiol. 2012 Oct;50(10):3159-67. doi: 10.1128/JCM.01251-12. Epub 2012 Jul 11. J Clin Microbiol. 2012. PMID: 22785194 Free PMC article.
References
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. - PubMed
-
- Arauz-Ruiz, P., H. Norder, B. H. Robertson, and L. O. Magnius. 2002. Genotype H: a new Amerindian genotype of hepatitis B virus revealed in Central America. J. Gen. Virol. 83:2059-2073. - PubMed
-
- Büchen-Osmond, C. (ed.). 2003. Hepadnaviridae. In ICTVdB—the Universal Virus Database, version 3. ICTVdB Management, Columbia University, New York, N.Y. [Online.] http://www.ncbi.nlm.nih.gov/ICTVdb/ICTVdB/30000000.htm.
-
- Burgener, M., U. Candrian, and M. Gilgen. 2003. Comparative evaluation of four large-volume RNA extraction kits in the isolation of viral RNA from water samples. J. Virol. Methods 108:165-170. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

