Recognition of T-rich single-stranded DNA by the cold shock protein Bs-CspB in solution
- PMID: 16956971
- PMCID: PMC1636342
- DOI: 10.1093/nar/gkl376
Recognition of T-rich single-stranded DNA by the cold shock protein Bs-CspB in solution
Abstract
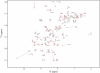
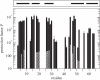
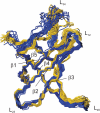
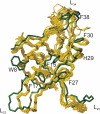
Cold shock proteins (CSP) belong to the family of single-stranded nucleic acid binding proteins with OB-fold. CSP are believed to function as 'RNA chaperones' and during anti-termination. We determined the solution structure of Bs-CspB bound to the single-stranded DNA (ssDNA) fragment heptathymidine (dT7) by NMR spectroscopy. Bs-CspB reveals an almost invariant conformation when bound to dT7 with only minor reorientations in loop beta1-beta2 and beta3-beta4 and of few aromatic side chains involved in base stacking. Binding studies of protein variants and mutated ssDNA demonstrated that Bs-CspB associates with ssDNA at almost diffusion controlled rates and low sequence specificity consistent with its biological function. A variation of the ssDNA affinity is accomplished solely by changes of the dissociation rate. 15N NMR relaxation and H/D exchange experiments revealed that binding of dT7 increases the stability of Bs-CspB and reduces the sub-nanosecond dynamics of the entire protein and especially of loop beta3-beta4.
Figures


References
-
- Graumann P.L., Marahiel M.A. A superfamily of proteins that contain the cold-shock domain. Trends Biochem. Sci. 1998;23:286–290. - PubMed
-
- Jones P.G., Inouye M. RbfA, a 30S ribosomal binding factor, is a cold-shock protein whose absence triggers the cold-shock response. Mol. Microbiol. 1996;21:1207–1218. - PubMed
-
- Phadtare S., Inouye M., Severinov K. The nucleic acid melting activity of Escherichia coli CspE is critical for transcription antitermination and cold acclimation of cells. J. Biol. Chem. 2002;277:7239–7245. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

