Sequence-specific recognition of colicin E5, a tRNA-targeting ribonuclease
- PMID: 16963495
- PMCID: PMC1635277
- DOI: 10.1093/nar/gkl629
Sequence-specific recognition of colicin E5, a tRNA-targeting ribonuclease
Abstract
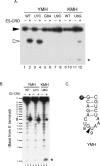
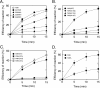
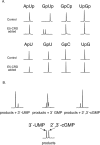
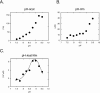
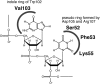
Colicin E5 is a novel Escherichia coli ribonuclease that specifically cleaves the anticodons of tRNA(Tyr), tRNA(His), tRNA(Asn) and tRNA(Asp). Since this activity is confined to its 115 amino acid long C-terminal domain (CRD), the recognition mechanism of E5-CRD is of great interest. The four tRNA substrates share the unique sequence UQU within their anticodon loops, and are cleaved between Q (modified base of G) and 3' U. Synthetic minihelix RNAs corresponding to the substrate tRNAs were completely susceptible to E5-CRD and were cleaved in the same manner as the authentic tRNAs. The specificity determinant for E5-CRD was YGUN at -1 to +3 of the 'anticodon'. The YGU is absolutely required and the extent of susceptibility of minihelices depends on N (third letter of the anticodon) in the order A > C > G > U accounting for the order of susceptibility tRNA(Tyr) > tRNA(Asp) > tRNA(His), tRNA(Asn). Contrastingly, we showed that GpUp is the minimal substrate strictly retaining specificity to E5-CRD. The effect of contiguous nucleotides is inconsistent between the loop and linear RNAs, suggesting that nucleotide extension on each side of GpUp introduces a structural constraint, which is reduced by a specific loop structure formation that includes a 5' pyrimidine and 3' A.
Figures
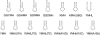
Similar articles
-
Structural basis for sequence-dependent recognition of colicin E5 tRNase by mimicking the mRNA-tRNA interaction.Nucleic Acids Res. 2006;34(21):6074-82. doi: 10.1093/nar/gkl729. Epub 2006 Nov 11. Nucleic Acids Res. 2006. PMID: 17099236 Free PMC article.
-
Structural and mutational studies of the catalytic domain of colicin E5: a tRNA-specific ribonuclease.Biochemistry. 2005 Aug 9;44(31):10494-500. doi: 10.1021/bi050749s. Biochemistry. 2005. PMID: 16060658
-
Colicin E5 ribonuclease domain cleaves Saccharomyces cerevisiae tRNAs leading to impairment of the cell growth.J Biochem. 2009 Apr;145(4):461-6. doi: 10.1093/jb/mvp004. Epub 2009 Jan 16. J Biochem. 2009. PMID: 19151102
-
tRNA-targeting ribonucleases: molecular mechanisms and insights into their physiological roles.Biosci Biotechnol Biochem. 2016 Jun;80(6):1037-45. doi: 10.1080/09168451.2016.1148579. Epub 2016 Mar 11. Biosci Biotechnol Biochem. 2016. PMID: 26967967 Review.
-
[The effect of modification of tRNA nucleotide-37 on the tRNA interaction with the P- and A-site of the 70S ribosome Escherichia coli].Mol Biol (Mosk). 2006 Jul-Aug;40(4):669-83. Mol Biol (Mosk). 2006. PMID: 16913226 Review. Russian.
Cited by
-
Phospho-RNA sequencing with circAID-p-seq.Nucleic Acids Res. 2022 Feb 28;50(4):e23. doi: 10.1093/nar/gkab1158. Nucleic Acids Res. 2022. PMID: 34850942 Free PMC article.
-
Development of a Flow Through-Based Limited Digestion Approach for High-Throughput and High-Sequence Coverage Mapping of Therapeutic mRNAs.Anal Chem. 2024 Oct 22;96(42):16994-17003. doi: 10.1021/acs.analchem.4c04384. Epub 2024 Oct 11. Anal Chem. 2024. PMID: 39391985 Free PMC article.
-
Growth-regulating Mycobacterium tuberculosis VapC-mt4 toxin is an isoacceptor-specific tRNase.Nat Commun. 2015 Jul 9;6:7480. doi: 10.1038/ncomms8480. Nat Commun. 2015. PMID: 26158745 Free PMC article.
-
The Versatile Roles of the tRNA Epitranscriptome during Cellular Responses to Toxic Exposures and Environmental Stress.Toxics. 2019 Mar 25;7(1):17. doi: 10.3390/toxics7010017. Toxics. 2019. PMID: 30934574 Free PMC article. Review.
-
Metatranscriptomic analysis of microbes in an Oceanfront deep-subsurface hot spring reveals novel small RNAs and type-specific tRNA degradation.Appl Environ Microbiol. 2012 Feb;78(4):1015-22. doi: 10.1128/AEM.06811-11. Epub 2011 Dec 9. Appl Environ Microbiol. 2012. PMID: 22156430 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

