Active TEM-1 beta-lactamase mutants with random peptides inserted in three contiguous surface loops
- PMID: 16963643
- PMCID: PMC2242396
- DOI: 10.1110/ps.062303606
Active TEM-1 beta-lactamase mutants with random peptides inserted in three contiguous surface loops
Abstract
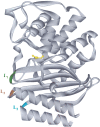
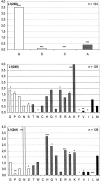
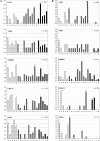
Engineering of alternative binding sites on the surface of an enzyme while preserving the enzymatic activity would offer new opportunities for controlling the activity by binding of non-natural ligands. Loops and turns are the natural substructures in which binding sites might be engineered with this purpose. We have genetically inserted random peptide sequences into three relatively rigid and contiguous loops of the TEM-1 beta-lactamase and assessed the tolerance to insertion by the percentage of active mutants. Our results indicate that tolerance to insertion could not be correlated to tolerance to mutagenesis. A turn between two beta-strands bordering the active site was observed to be tolerant to random mutagenesis but not to insertions. Two rigid loops comprising rather well-conserved amino acid residues tolerated insertions, although with some constraints. Insertions between the N-terminal helix and the first beta-strand generated active libraries if cysteine residues were included at both ends of the insert, suggesting the requirement for a stabilizing disulfide bridge. Random sequences were relatively well accommodated within the loop connecting the final beta-strand to the C-terminal helix, particularly if the wild-type residue was retained at one of the loops' end. This suggests two strategies for increasing the percentage of active mutants in insertion libraries. The amino acid distribution in the engineered loops was analyzed and found to be less biased against hydrophobic residues than in natural medium-sized loops. The combination of these activity-selected libraries generated a huge library containing active hybrid enzymes with all three loops modified.
Figures

Similar articles
-
An engineered disulfide bond between residues 69 and 238 in extended-spectrum beta-lactamase Toho-1 reduces its activity toward third-generation cephalosporins.Biochemistry. 2004 Dec 21;43(50):15737-45. doi: 10.1021/bi048488u. Biochemistry. 2004. PMID: 15595829
-
Engineering allosteric regulation into the hinge region of a circularly permuted TEM-1 beta-lactamase.Protein Eng Des Sel. 2010 Sep;23(9):699-709. doi: 10.1093/protein/gzq041. Epub 2010 Jun 30. Protein Eng Des Sel. 2010. PMID: 20591901
-
Probing beta-lactamase structure and function using random replacement mutagenesis.Proteins. 1992 Sep;14(1):29-44. doi: 10.1002/prot.340140106. Proteins. 1992. PMID: 1329081
-
[A turning point in the knowledge of the structure-function-activity relations of elastin].J Soc Biol. 2001;195(2):181-93. J Soc Biol. 2001. PMID: 11727705 Review. French.
-
'Anticalins': a new class of engineered ligand-binding proteins with antibody-like properties.J Biotechnol. 2001 Jun;74(4):257-75. doi: 10.1016/s1389-0352(01)00020-4. J Biotechnol. 2001. PMID: 11526907 Review.
Cited by
-
Ligand binding and allostery can emerge simultaneously.Protein Sci. 2007 May;16(5):929-37. doi: 10.1110/ps.062706007. Epub 2007 Mar 30. Protein Sci. 2007. PMID: 17400921 Free PMC article.
-
Fitness Effects of Single Amino Acid Insertions and Deletions in TEM-1 β-Lactamase.J Mol Biol. 2019 May 31;431(12):2320-2330. doi: 10.1016/j.jmb.2019.04.030. Epub 2019 Apr 26. J Mol Biol. 2019. PMID: 31034887 Free PMC article.
-
The Bacillus licheniformis BlaP beta-lactamase as a model protein scaffold to study the insertion of protein fragments.Protein Sci. 2007 Oct;16(10):2260-71. doi: 10.1110/ps.072912407. Protein Sci. 2007. PMID: 17893363 Free PMC article.
-
Selection of allosteric beta-lactamase mutants featuring an activity regulation by transition metal ions.Protein Sci. 2006 Oct;15(10):2335-43. doi: 10.1110/ps.062304406. Epub 2006 Sep 8. Protein Sci. 2006. PMID: 16963642 Free PMC article.
-
Deep indel mutagenesis reveals the impact of amino acid insertions and deletions on protein stability and function.Nat Commun. 2025 Mar 17;16(1):2617. doi: 10.1038/s41467-025-57510-5. Nat Commun. 2025. PMID: 40097423 Free PMC article.
References
-
- Batori, V., Koide, A., Koide, S. 2002. Exploring the potential of the monobody scaffold: Effects of loop elongation on the stability of a fibronectin type III domain. Protein Eng. 15: 1015–1020. - PubMed
-
- Binz, K.P., Amstutz, P., Plückthun, A. 2005. Engineering novel binding proteins from nonimmunoglobulin domains. Nat. Biotechnol. 23: 1257–1268. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

