Amyloid formation in denatured single-mutant lysozymes where residual structures are modulated
- PMID: 16963644
- PMCID: PMC2242399
- DOI: 10.1110/ps.062258206
Amyloid formation in denatured single-mutant lysozymes where residual structures are modulated
Abstract
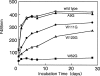
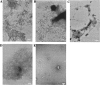
Reduced hen lysozyme has a residual structure involving long-range interaction. It has been demonstrated that a single mutation (A9G, W62G, W111G, or W123G) in the residual structure differently modulates the long-range interactions of reduced lysozyme. To examine whether such variations in the residual structure affect amyloid formation, reduced and alkylated mutant lysozymes were incubated under the amyloid-fibrillation condition. From the analyses of CD spectra and thioflavine T fluorescences, it was suggested that variation in residual structure led to different amyloid formation. Interestingly, the extent of amyloid formation did not always correlate with the extent to which the residual structure was maintained, resulting in the involvement of a hydrophobic cluster normally contained in W111 in the reduced lysozyme.
Figures


Similar articles
-
A particular hydrophobic cluster in the residual structure of reduced lysozyme drastically affects the amyloid fibrils formation.Biochem Biophys Res Commun. 2007 May 11;356(3):769-72. doi: 10.1016/j.bbrc.2007.03.043. Epub 2007 Mar 16. Biochem Biophys Res Commun. 2007. PMID: 17382294
-
Effect of the structure of the denatured state of lysozyme on the aggregation reaction at the early stages of folding from the reduced form.J Mol Biol. 2005 Mar 18;347(1):159-68. doi: 10.1016/j.jmb.2005.01.022. Epub 2005 Jan 21. J Mol Biol. 2005. PMID: 15733925
-
Amyloid fibril formation of hen lysozyme depends on the instability of the C-helix (88-99).Biosci Biotechnol Biochem. 2008 Jun;72(6):1523-30. doi: 10.1271/bbb.80032. Epub 2008 Jun 7. Biosci Biotechnol Biochem. 2008. PMID: 18540107
-
Amide hydrogen exchange in a highly denatured state. Hen egg-white lysozyme in urea.J Mol Biol. 1994 Apr 1;237(3):247-54. doi: 10.1006/jmbi.1994.1228. J Mol Biol. 1994. PMID: 8145239 Review.
-
From the globular to the fibrous state: protein structure and structural conversion in amyloid formation.Q Rev Biophys. 1998 Feb;31(1):1-39. doi: 10.1017/s0033583598003400. Q Rev Biophys. 1998. PMID: 9717197 Review. No abstract available.
Cited by
-
Mechanism for retardation of amyloid fibril formation by sugars in Vλ6 protein.Protein Sci. 2013 Apr;22(4):467-74. doi: 10.1002/pro.2228. Epub 2013 Feb 21. Protein Sci. 2013. PMID: 23389799 Free PMC article.
-
Folding versus aggregation: polypeptide conformations on competing pathways.Arch Biochem Biophys. 2008 Jan 1;469(1):100-17. doi: 10.1016/j.abb.2007.05.015. Epub 2007 Jun 8. Arch Biochem Biophys. 2008. PMID: 17588526 Free PMC article. Review.
References
-
- Bellotti, V., Mangione, P., Merlini, G. 2000. Review: Immunoglobulin light chain amyloidosis—The archetype of structural and pathogenic variability. J. Struct. Biol. 130: 280–289. - PubMed
-
- Chakraborty, C., Nandi, S., Jana, S. 2005. Prion disease: A deadly disease for protein misfolding. Curr. Pharm. Biotechnol. 6: 167–177. - PubMed
-
- Damaschun, G., Damaschun, H., Fabian, H., Gast, K., Krober, R., Wieske, M., Zirwer, D. 2000. Conversion of yeast phosphoglycerate kinase into amyloid-like structure. Proteins 39: 204–211. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

