Exportin-5 orthologues are functionally divergent among species
- PMID: 16963774
- PMCID: PMC1635293
- DOI: 10.1093/nar/gkl663
Exportin-5 orthologues are functionally divergent among species
Abstract
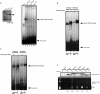
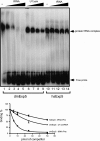
Exportin-5, an evolutionarily conserved nuclear export factor belonging to the importin-beta family of proteins, is known to play a role in the nuclear export of small noncoding RNAs such as precursors of microRNA, viral minihelix RNA and a subset of tRNAs in mammalian cells. In this study, we show that the exportin-5 orthologues from different species such as human, fruit fly and yeast exhibit diverged functions. We found that Msn5p, a yeast exportin-5 orthologue, binds double-stranded RNAs and that it prefers a shorter 22 nt, double-stranded RNA to approximately 80 nt pre-miRNA, even though both of these RNAs share a similar terminal structure. Furthermore, we found that Drosophila exportin-5 binds pre-miRNAs and that amongst the exportin-5 orthologues tested, it shows the highest affinity for tRNAs. The knockdown of Drosophila exportin-5 in cultured cells decreased the amounts of tRNA as well as miRNA, whereas the knock down of human exportin-5 in cultured cells affected only miRNA but not tRNA levels. These results indicate that double-stranded RNA binding ability is an inherited functional characteristic of the exportin-5 orthologues and that Drosophila exportin-5 functions as an exporter of tRNAs as well as pre-miRNAs in the fruit fly that lacks the orthologous gene for exportin-t.
Figures


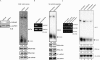

Similar articles
-
Exportin-5 binding precedes 5'- and 3'-end processing of tRNA precursors in Drosophila.J Biol Chem. 2024 Sep;300(9):107632. doi: 10.1016/j.jbc.2024.107632. Epub 2024 Aug 2. J Biol Chem. 2024. PMID: 39098529 Free PMC article.
-
Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs.RNA. 2004 Feb;10(2):185-91. doi: 10.1261/rna.5167604. RNA. 2004. PMID: 14730017 Free PMC article.
-
Selective nuclear export mechanism of small RNAs.Curr Opin Struct Biol. 2011 Feb;21(1):101-8. doi: 10.1016/j.sbi.2010.11.004. Epub 2010 Dec 7. Curr Opin Struct Biol. 2011. PMID: 21145229 Review.
-
Minihelix-containing RNAs mediate exportin-5-dependent nuclear export of the double-stranded RNA-binding protein ILF3.J Biol Chem. 2004 Jan 9;279(2):884-91. doi: 10.1074/jbc.M306808200. Epub 2003 Oct 21. J Biol Chem. 2004. PMID: 14570900
-
Exportin t and Exportin 5: tRNA and miRNA biogenesis - and beyond.Biol Chem. 2012 Jul;393(7):599-604. doi: 10.1515/hsz-2012-0146. Biol Chem. 2012. PMID: 22944664 Review.
Cited by
-
Recent advances in the functional explorations of nuclear microRNAs.Front Immunol. 2023 Feb 22;14:1097491. doi: 10.3389/fimmu.2023.1097491. eCollection 2023. Front Immunol. 2023. PMID: 36911728 Free PMC article. Review.
-
tRNA biology charges to the front.Genes Dev. 2010 Sep 1;24(17):1832-60. doi: 10.1101/gad.1956510. Genes Dev. 2010. PMID: 20810645 Free PMC article. Review.
-
Genetic and epigenetic association studies suggest a role of microRNA biogenesis gene exportin-5 (XPO5) in breast tumorigenesis.Int J Mol Epidemiol Genet. 2011 Jan 1;2(1):9-18. Epub 2010 Nov 25. Int J Mol Epidemiol Genet. 2011. PMID: 21552306 Free PMC article.
-
MicroRNA expression and its association with DNA repair in preimplantation embryos.J Reprod Dev. 2016 Jun 17;62(3):225-34. doi: 10.1262/jrd.2015-167. Epub 2016 Feb 5. J Reprod Dev. 2016. PMID: 26853522 Free PMC article. Review.
-
Three tRNA nuclear exporters in S. cerevisiae: parallel pathways, preferences, and precision.Nucleic Acids Res. 2022 Sep 23;50(17):10140-10152. doi: 10.1093/nar/gkac754. Nucleic Acids Res. 2022. PMID: 36099418 Free PMC article.
References
-
- Chook Y.M., Blobel G. Karyopherins and nuclear import. Curr. Opin. Struct. Biol. 2001;11:703–715. - PubMed
-
- Kuersten S., Ohno M., Mattaj I.W. Nucleocytoplasmic transport: Ran, beta and beyond. Trends Cell Biol. 2001;11:497–503. - PubMed
-
- Weis K. Nucleocytoplasmic transport: cargo trafficking across the border. Curr. Opin. Cell Biol. 2002;14:328–335. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

